Glycosylation Analysis Service for Influenza Virus Glycoprotein
Roles of Viral Glycoproteins on Influenza Virus Evolution
Influenza viruses cause both seasonal epidemics and occasional pandemics in humans. The two main envelope glycoproteins on the surface of influenza viruses are hemagglutinin (HA) and neuraminidase (NA), both undergoing N-linked glycosylation. HA and NA glycoproteins play a crucial role in the recognition of host immune cells and are considered the primary antigenic determinants of the virus. Influenza A viruses, the greatest concern for public health, are categorized into various subtypes according to variations in their surface glycoproteins, specifically 18 HA (H1-H18) and 11 NA (N1-N11) subtypes. Glycosylation profile changes in viral glycoproteins, particularly HA, have a significant impact on the evolution of influenza A viruses. HA antigenic drift, which involves modifications in the number and location of glycosylation sites over time, can lead to alterations in various biological activities and the antigenic properties of the virus strain. This allows the novel virus subtypes to evade neutralizing antibodies and avoid immune recognition by host cells. Given the pivotal role of glycans in determining antigenicity, it is crucial to explore the glycosylation patterns of influenza viruses to improve the development of vaccines and antiviral drugs.
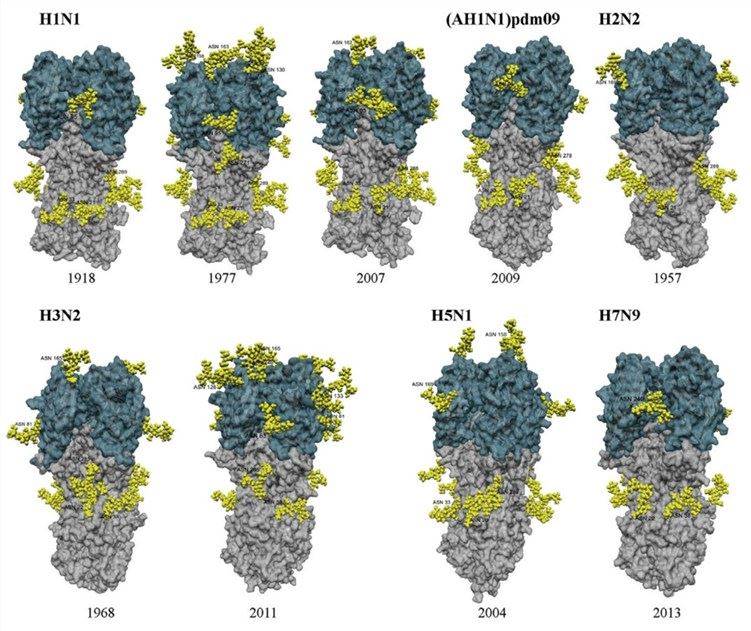 Fig.1 Structural models of the HA from different influenza A virus subtypes.1
Fig.1 Structural models of the HA from different influenza A virus subtypes.1
Glycosylation Analysis Services for Influenza Virus Glycoprotein at Creative Biolabs
The glycosylation patterns on these glycoproteins play a crucial role in the evolution of influenza viruses, understanding these glycosylation patterns is essential for improving influenza vaccines and antiviral strategies. Creative Biolabs has developed glycosylation analysis services for viral glycoproteins, specifically focusing on influenza virus glycoproteins.
Based on an in-house developed server, we provide sequence-specific prediction of potential glycosylation sites by analyzing adjacent primary sequences within the viral glycoproteins. For further confirmation, we utilize high-resolution HCD/CID-MS/MS tandem mass spectrometry to thoroughly analyze the glycosylation profiles of HA and NA glycoproteins from various influenza A virus subtypes, such as H1N1, H3N2, H5N1, H7N9. By combining predictive analysis with experimental validation, we ensure accurate and thorough glycosylation analysis for influenza virus glycoproteins, which provides valuable insights into how glycans and glycosylation patterns impact the biological activity of these glycoproteins and the evolution of influenza viruses.
In-depth Characterization of Viral Glycosylation at Creative Biolabs
Through comprehensive characterization, we provide essential glycosylation information on viral glycoproteins, including:
-
Identifying N-glycan structures and assessing glycan heterogeneity: We determine the specific types of N-glycan structures and analyze the diversity of glycan compositions attached to each glycosylation site. Research has indicated the presence of substantial glycan microheterogeneity in HA and NA glycoproteins. This microheterogeneity is closely linked to the biological activities and immune recognition of influenza viruses.
-
Evaluating glycosylation site localization and occupancy: We perform in-depth analyses of the changes in glycosylation site localization, site occupancy, and conserved sites among different influenza virus subtypes. It has been reported that HA possesses high glycosylation site occupancy and the number of potential glycosylation sites in both HA and NA of seasonal strains has increased throughout virus evolution. Distinct N-glycosylation sites and an increase in the number of glycosylation sites have been linked to immune evasion and attenuation.
Advantages of Our Services
-
High-resolution and accuracy detection of tandem mass spectrometry
-
Comprehensive analysis encompassing glycosylation sites and glycan structures
-
Glycosylation analysis for HA and NA glycoproteins from all influenza virus subtypes
-
Systematic comparison of glycosylation profiles among different virus subtypes
-
Professional team experienced in glycoprotein analysis
Published data
Glycosylation is a critical quality indicator in the production of recombinant biopharmaceuticals. Here, an innovative sugar analysis process was proposed, using two orthogonal LC-MS methods to perform comprehensive sugar structure and site-specific analysis of glycopeptides of viral antigens. The research focused on the N-glycosylation characteristics of neuraminidase (NA) and hemagglutinin (HA) in influenza A virus (IAV). By comparing IAVs cultured in different Madin-Darby canine kidney (MDCK) cell lines, it was found that the two proteins showed similar N-glycosylation patterns. Still, there were differences in the relative abundance of specific sugar structures. The developed mass spectrometry analysis workflow enables site-specific glycosylation analysis, which provides new ideas for the quality assessment of viral antigens in future vaccine development.
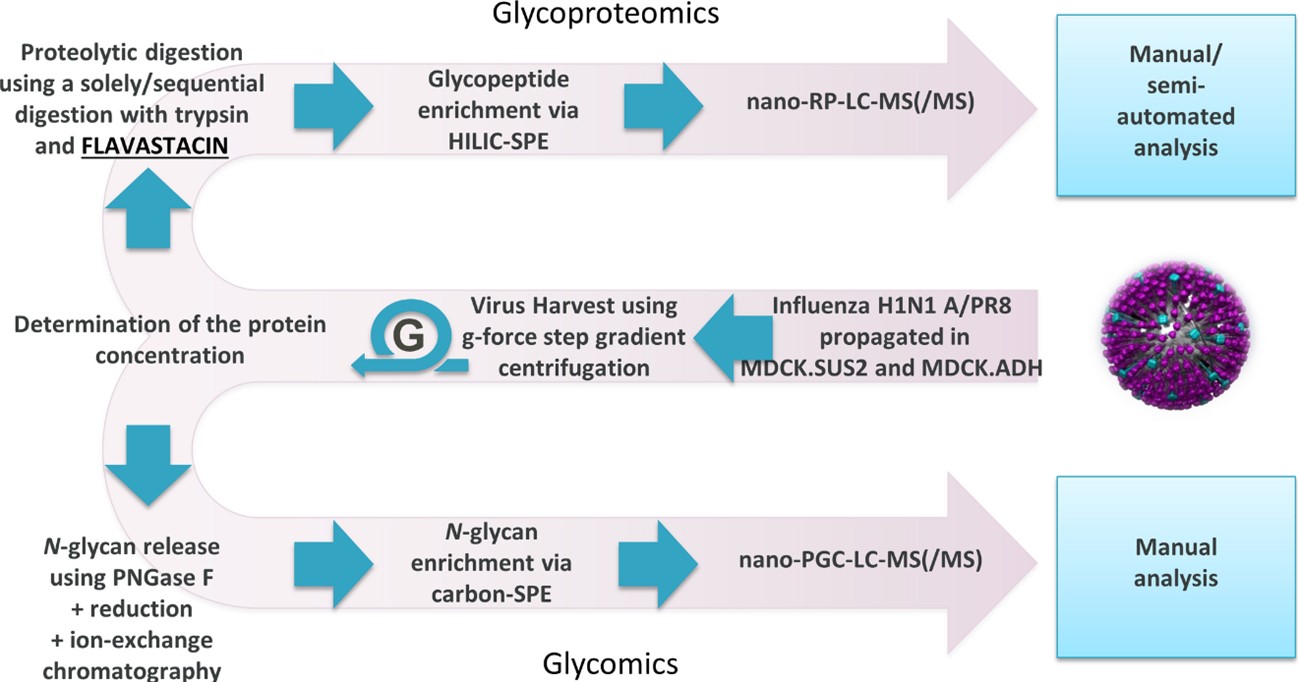 Fig.2 Glycosylation analysis process of IAV glycoproteins.2
Fig.2 Glycosylation analysis process of IAV glycoproteins.2
Creative Biolabs has developed comprehensive glycosylation analysis services for influenza virus glycoproteins to investigate the impacts of glycosylation patterns of HA and NA glycoprotein on influenza virus evolution and advance the development of improved vaccines. For further details or specific inquiries about our influenza virus glycoprotein analysis services, please don't hesitate to contact us.
FAQ
Q1: How do you identify potential glycosylation sites on influenza virus glycoproteins?
A1: We utilize an in-house developed server to predict potential N-glycosylation sites and employ high-resolution mass spectrometry techniques to analyze the glycosylation profiles of viral glycoproteins. We combine predictive with experimental results and provide an accurate assessment of glycosylation on influenza virus glycoproteins.
Q2: How do you analyze and determine the diversity of N-glycan structures?
A2: Our team specializes in the study of glycosylation and its impact on biological functions, backed by years of relevant experience. We utilize mass spectrometry to identify various N-glycan structures and assess their compositional diversity. This approach helps reveal the microheterogeneity of HA and NA and its implications for immune recognition.
Q3: How can I obtain a quote and timeline for this service?
A3: Please contact us in time and we will provide a detailed quote and time required according to your specific needs.
Customer Review
High Analysis Accuracy
“Our research team was very satisfied with the results of the viral glycosylation analysis provided by Creative Biolabs. The data obtained by high-resolution mass spectrometry technology was extremely accurate, which provided a solid foundation for our subsequent research.”
Flexibility And Comprehensive Data
“According to the needs of our different experimental stages, Creative Biolabs flexibly adjusted the service content to make the analysis more in line with our research goals. The glycosylation site and N-sugar structure analysis they provided was very detailed, allowing us to fully understand the characteristics of influenza virus glycoproteins.”
References
-
Tate, Michelle D., et al. "Playing hide and seek: how glycosylation of the influenza virus hemagglutinin can modulate the immune response to infection." Viruses 6.3 (2014): 1294-1316. Distributed under Open Access license CC BY 3.0, without modification.
-
Pralow, Alexander, et al. "Comprehensive N‐glycosylation analysis of the influenza A virus proteins HA and NA from adherent and suspension MDCK cells." The FEBS Journal 288.16 (2021): 4869-4891. Distributed under Open Access license CC BY 4.0, without modification.
For Research Use Only.
Related Services
 Fig.1 Structural models of the HA from different influenza A virus subtypes.1
Fig.1 Structural models of the HA from different influenza A virus subtypes.1
 Fig.2 Glycosylation analysis process of IAV glycoproteins.2
Fig.2 Glycosylation analysis process of IAV glycoproteins.2