Gut Microbiota Metaproteomic Service
Proteomics, initially defined as the study of all proteins expressed by a single organism, is gradually applied to complex bacterial communities. Thus, the analysis of the protein content of the microbial communities, such as gut microbiota is now named “metaproteomics”. Creative Biolabs offers gut microbiota metaproteomic service to identify broad protein profiles in complex samples, such as gut microbiota.
Human Gut Microbiome
The human gut microbiota consists of billions of microorganisms which are ten times more numerous than human cells. These microorganisms include bacteria, viruses, archaea, yeasts, and protozoa. Under normal conditions, gut microbiota lives in mutual coexistence with the body and has a considerable impact on human health and physiology. The microbial imbalance, also known as dysbiosis, disrupts the microbiota composition and can lead to intestinal permeability. Dysbiosis can occur due to several factors, such as environment, aging, diet, and the immune system. As a result, it has been associated with inflammatory bowel diseases, obesity, metabolic, immune, and neurological diseases, and cancers. Metaproteomics, a tool for the large-scale analysis of proteins in microbiomes, allows researchers to address a diversity of questions related to functions and interactions in microbiomes.
Workflow of Metaproteomic Analysis
Our metaproteomic analysis typically comprises the following 4 steps:
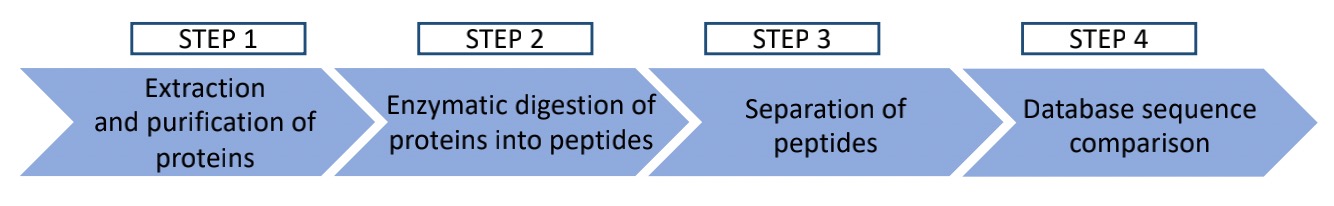 Fig.1 General workflow of metaproteomic analysis.
Fig.1 General workflow of metaproteomic analysis.
Our Service
-
Stool Sample Preparation
Fecal samples are primarily performed in the study of the metaproteome of gut microbiota. However, there is a complicated environmental matrix exited in stool which can interfere with protein characterization studies. The challenges faced in fecal samples include 1) a mix of gram-positive and gram-negative cells with various envelopes structures. 2) an abundance of host proteins. 3) the presence of proteins derived from consumed and undigested foods. 4) various physico-chemical properties of proteins involved in their solubility.
Creative Biolabs offers comprehensive steps during stool sample preparation for the metaproteomic characterization of the human gut microbiota:
-
General differential centrifugation and double filtering (DF) differential separation
-
Enzymatically digested (trypsin)
-
Pre-concentration steps, such as filter-aided sample preparation (FASP)
-
Removal of detergents and salts
Except for fecal, the sample of colonic lavage and colon biopsies are also employed in the metaproteomic analysis of the gut microbiome.
-
Pre-fractionation and Mass Spectrometry
Creative Biolabs offers gel electrophoresis to separate proteins and liquid chromatography to separate peptides:
-
Two-dimensional polyacrylamide gel electrophoresis (2D-PAGE) for protein separation
-
High-performance liquid chromatography (HPLC) and reverse phase (RP) liquid chromatography (LC) for peptide separation
Creative Biolabs offers MS/MS to monitor the mass of the peptides and their induced fragments.
-
Metaproteomics Data Computation
The processing of metagenomic data must be carried out carefully to ensure the best quality of the resulting metaproteomic databases. Creative Biolabs offers the following database to identify proteins.
-
Conventional sequence database search
-
Customized iterative database approach
-
De novo sequencing search
Applications of Gut Microbiota Metaproteomic
-
Providing information about microbiomes, including composition, abundance dynamics, the metabolism and physiology of individual members, and the host.
-
Providing gene expression responses to changing conditions, localization of potential host–microbe interaction proteins, and the uptake of substrates labeled with heavy carbon or nitrogen.
-
Characterizing the functional roles of gut microbiota in health and disease conditions, unraveling the molecular mechanism underlying homeostasis and disease pathogenesis.
-
Biomarker development to discriminate between normal individuals and diseased (i.e., inflammatory bowel disease or colon cancer) patients as well as to monitor disease activity and prognosis.
If you are interested in our gut microbiota metaproteomic service, please feel free to contact us.
For Research Use Only | Not For Clinical Use


 Fig.1 General workflow of metaproteomic analysis.
Fig.1 General workflow of metaproteomic analysis.
 Download our brochure
Download our brochure