Gut Microbiota Metatranscriptomic Service
Metatranscriptomics is employed to determine microbiota gene expression and regulation. Creative Biolabs offers comprehensive gut microbiota metatranscriptomic services to help you identify, classify, and quantity the gut microbiota.
Gut Microbiome
Gut microbiome consists of more than 1,000 bacteria species. The Firmicutes, Bacteriodetes, Actinobacteria, and Proteobacteria are the most common phyla. They play an important multifactorial role in human physiology. They are important in controlling pathogen colonization and in immune system development, and they help us in our digestion by hydrolyzing the compounds in our diet. Besides, changes in microbiota populations, also known as dysbiosis, have been associated with a number of human conditions, including obesity, inflammatory bowel disease, and nonalcoholic fatty liver disease.
Characteristics of Metatranscriptome
Metatranscriptome analysis could demonstrate the functions of a potential repertoire of bacteria. This information is useful in elucidating accurately which of the genes are transcribed and to what extent. Furthermore, functional data show active metabolic pathways in the bacterial communities and can be associated with particular environmental conditions. Compared to metagenomics, metatranscriptomics offers more information about transcriptionally active populations.
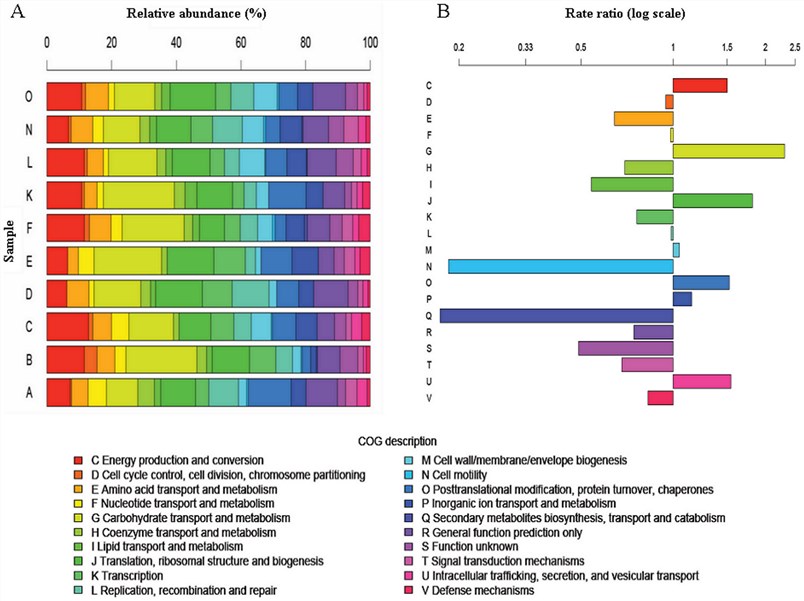 Fig.1 Metatranscriptomic approach to analyze the functional human gut microbiota.1
Fig.1 Metatranscriptomic approach to analyze the functional human gut microbiota.1
Applications of Metatranscriptomic
-
Microbial activity assessment
-
Microbiome antisense RNA study
-
Microbiome–immune interactions assessment
-
Microbiome small noncoding RNAs study
Our Services
-
Isolation and Processing of Microbiome mRNA
-
Data Analysis of Metatranscriptomic
Creative Biolabs provides one-stop metatranscriptomic analysis services including:
-
Non-mRNA reads
-
Host reads
-
Filtering and trimming low-quality reads
-
Quality control
-
Identifying the open-reading frames
-
Mapping the reads to a reference database
-
Normalization
-
Calculations of the gene expression levels
Advantages of Gut Microbiota Metatranscriptomic Service
Metatranscriptomic sequences all genetic material in the sample.
Metatranscriptomic does not need cloning the DNA before sequencing, which removes one of the main biases and bottlenecks in environmental sampling.
Metatranscriptomic shows all species diversity, including eukarya, archaea, bacteria and viruses.
With the price of next-generation sequencing (NGS) continues falling, metatranscriptomics allows microbial ecology to be investigated at a much greater scale and detail.
Creative Biolabs offers metatranscriptome sequencing services to help you gain important insights into many functions such as immune response, disease progression, and antibiotic resistance. We also provide nongenomic-based approaches which can complement metagenomics and metatranscriptomics data to improve the understanding of complex pathways in the dynamic environment of microbiota communities. If you are interested in our metatranscriptomic service, please feel free to contact us.
Reference
-
Gosalbes, María José, et al. "Metatranscriptomic approach to analyze the functional human gut microbiota." PloS one 6.3 (2011): e17447. Distributed under Open Access license CC BY 4.0, without modification.
For Research Use Only | Not For Clinical Use


 Fig.1 Metatranscriptomic approach to analyze the functional human gut microbiota.1
Fig.1 Metatranscriptomic approach to analyze the functional human gut microbiota.1
 Download our brochure
Download our brochure