By integrating next-generation sequencing (NGS) data with flow cytometry data, Creative Biolabs is able to provide the service of full-length Ig heavy (IgH) and light (Igκ/IgL) sequence analysis by B cell receptor (BCR) sequencing at the single-cell level, permitting the accurate correction of sequencing errors. The strategy is fully compatible with established procedures for antibody cloning and expression, and it can be used to study B cell repertoires based on Ig reactivity and function.
The Application of NGS for Full-length Ig Analysis
Antibodies are B cell antigen receptors encoded by two independent IgH and Igκ/IgL chain genes. Therefore, the accurate assessment of antibody repertoires and clonal relationships between individual B cells requires the combined analysis of IgH and Igκ/IgL genes from the same cell. Conventional single-cell PCR (scPCR) based antibody repertoire analysis uses Sanger sequencing of IgH and corresponding Igκ/IgL PCR products to assess antibody repertoires in mice and humans. It is a powerful tool to measure the diversity of antibody repertoires, allowing the functional assessment of B cell responses. However, it is not high throughput compatible, and scPCR is costly, time-consuming and limited to a relatively small number of several hundred input cells. With the advent of NGS technologies, high-throughput sequencing of IgH and Igκ/IgL light chains from a bulk of B cells has been widely used to analyze Ig gene repertoires in mice and humans. A major drawback of NGS from bulk cells is the inherent loss of information about the association between IgH and Igκ/IgL genes at the cellular level, which prevents the faithful functional assessment of antibody reactivity after Ig gene cloning and expression of recombinant mAbs in vitro. Physical linkage of IgH and Igκ/IgL PCR amplicons before sequencing can overcome this shortcoming but does not provide full-length Ig gene sequence information due to short NGS read lengths, and it is not compatible with the direct and high-throughput generation of recombinant mAbs. In addition, NGS is associated with high error rates. Computational filters can mask sequencing errors but cannot distinguish them from naturally occurring somatic mutations in Ig genes.
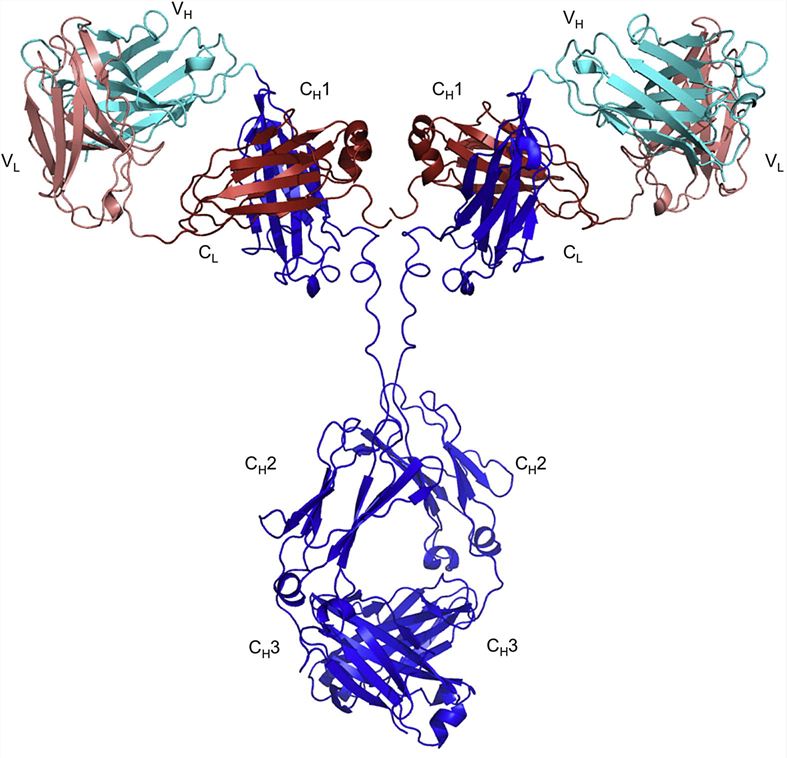 Fig.1 Structure of the human IgG molecule (Rouet et al. 2014).
Fig.1 Structure of the human IgG molecule (Rouet et al. 2014).
Full-length Ig Analysis at the Single-cell Level in Creative Biolabs
Combining IgH and Igκ/IgL chain gene PCR with NGS, Creative Biolabs has developed a two-dimensional barcoded primer matrix at the single-cell level. We combined the scPCR-based amplification of IgH and Igκ/IgL transcripts and NGS into a unified pipeline, which allows thousands of single B cells. Our Magic™ sequencing platform is scalable and can provide high-quality sequence information. The operation process roughly consists of (1) tissue preparation and single-cell sorting, (2) design of Tag and primer matrix, (3) single-cell PCR, and (4) sequencing on our Magic™ platform. Our scientist will analyze the sequences by our special sequence analysis pipeline. Importantly, the tag matrix can be used universally to barcode primer sets for the single-cell based amplification of Ig genes from any species.
Key Advantages
With years of research and development experience in the field of immunology, scientists of Creative Biolabs could provide full-length IgH and Igκ/IgL sequence analysis by BCR sequencing at the single-cell level. We are pleased to offer the best service with the most accurate results for our global customers.
Please contact us for more information and a detailed quote.
References
All listed services and products are For Research Use Only. Do Not use in any diagnostic or therapeutic applications.