PreciAbTM
PreciAb™ Platform
Antibody Design
Antibody Structure Modelling
Antibody-Antigen Complex Analysis
Computer-aided Affnity Maturation
Antibody Structure Determination
Antibody Reformatting
Antibody to ScFv Reformat
Antibody to VHH Reformat
Antibody Developability Prediction
Antibody Aggregation Prediction
Antibody Immunongenicity Prediction
Antibody Screening & Design
Company
About Us
Contact Us
Downloads
Careers
Events

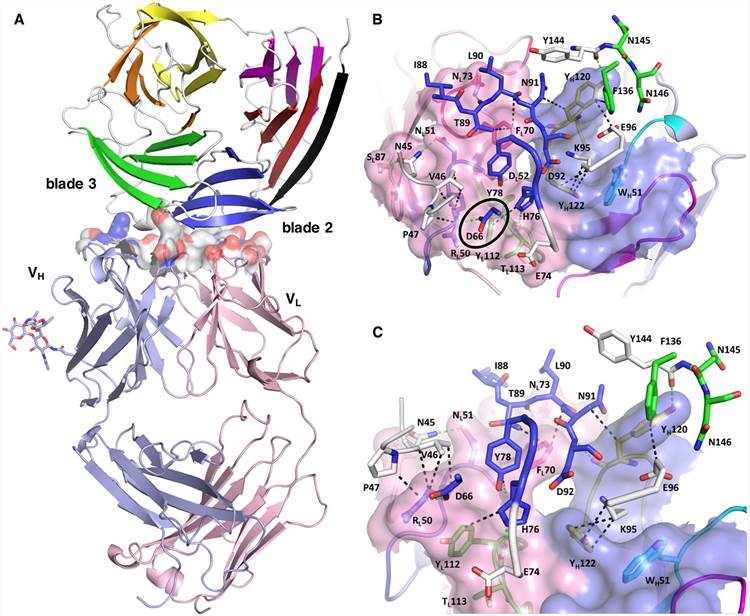 Fig.1 Structure of the PfCyRPA/c12 complex. Part A exhibits that the major interface is made by interactions between the light chain of c12 and blade 2 of PfCyRPA. Part A exhibits the details of the interface viewed from top onto the CDR loops.1
Fig.1 Structure of the PfCyRPA/c12 complex. Part A exhibits that the major interface is made by interactions between the light chain of c12 and blade 2 of PfCyRPA. Part A exhibits the details of the interface viewed from top onto the CDR loops.1

 More than 10 years of exploration and expansion
More than 10 years of exploration and expansion


