Phage display has proven to be a powerful technique for the interrogation of libraries containing millions or even billions of different peptides or proteins. Phage display libraries permit the selection of peptides and proteins, including antibodies, with high affinity and specificity for almost any target. Antibody phage display libraries obviate the need for immunization and the concomitant laborious hybridoma protocols for obtaining mAbs, directly afford the cloned antibody genes in single-chain Fv (scFv) or Fab format for convenient manipulation, and, importantly, can be derived from the human antibody repertoire. For more than a decade, phage displayed combinatorial antibody libraries have been used to generate and isolate a wide variety of antibodies. Combinatorial antibody library technology represents a powerful tool for discovering and designing antibodies that bind targets with high affinity and specificity. A crucial advantage of this technology is the direct link that exists between the experimental phenotype and its encapsulated genotype, which allows the evolution of the selected binders into optimized molecules. Using this technique, antibody genes have been cloned from multiple species or expressed directly from large man-made repertoires of antibody-encoding genes. Recent studies demonstrate that the technique allows for the in vitro evolution of antibodies to create molecules whose affinity for antigen exceeds that observed in nature.
The type of combinatorial antibody library
There are four types of antibody library in combinatorial antibody libraries: immune libraries, naïve, semisynthetic and synthetic libraries, the latter three have been included as “single-pot” libraries, as they are designed to isolate antibody fragments binding to every possible antigen in theory. We discriminate ‘naïve’ and ‘synthetic’ antibody libraries, depending on the source of immunoglobulin genes. For most applications, the availability of large pre-made collections of non-immune repertoires has superseded the use of immune repertoires.
Immune libraries from human or animals are constructed from immunized donors or natural infections and typically used in biopharmaceutical research to obtain an antibody against a particular target antigen. From immune libraries, V genes of these libraries contain hypermutations and are affinity matured, antibody fragments with monovalent dissociation constants in the nanomolar range can be isolated. Immune libraries are typically created and used in medical research to isolate an antibody fragment against one particular antigen, and therefore would not be the source of choice for the selection of a large number of different specificities.
The naïve libraries are construct from natural unimmunized human rearranged V genes, synthetic human V genes, or shuffled V genes. These universal (non-immunized) libraries differ from libraries derived from immune antibodies, in that the former are antigen-independent (i.e. a single library can be panned for specificity against virtually any antigen). Examples of naïve library type are the naïve human Fab library constructed by de Haard and colleagues, the McCafferty libraries, or the HAL scFv libraries. The affinity of Abs selected from a naïve library is proportional to the size of the library, ranging from 106–7 M–1 for a small library of 3×107 clones to 108–10 M–1 for a large repertoire of 1010 clones made by brute-force cloning.
Semi-synthetic libraries is constructed wherein diversity is controlled by the oligonucleotide synthesis. Semi-synthetic libraries are derived from unrearranged V-genes from pre-B cells or a single antibody framework with at least one complementary determining region (CDR) region genetically randomized. The combinations create new diversity. The character of the naïve repertoire will be influenced or edited by the animal and will be extremely susceptible to contamination from materials derived from activated B cells or plasma cells, which would limit the diversity of the library. Semi-synthetic libraries can overcome this diversity problem.
Synthetic libraries contain a framework with randomly integrated codons into CDRs. Antibodies are built artificially, by in vitro assembly of V-gene segments and D:J segments. V genes may be assembled by introducing a predetermined level of randomization of CDR regions into germline V-gene segments, or rearranged V-genes. Through synthetic library construction, new diversity can be obtained. Synthetic libraries are antigen-independent and particularly useful for unbiased selection of antibodies against any target antigen. These kinds of libraries were optimized in the past years by mimicking the in vivo amino acid distribution in the CDRs. Some of these libraries have yielded antibodies against many different antigens, including haptens, proteins, and cell-surface markers, but their affinities are typically in the micromolar range.
The construction of phage antibody Libraries
Library construction is facilitated by the ready availability of phagemid vectors, which allow for construction and display of libraries of all of these types of antibody fragments using a single rare cutting restriction enzyme. Selection of interesting members from the library based on the displayed antibodies’ binding specificity and affinity is generally performed over several rounds of selection and amplification in a process known as panning.
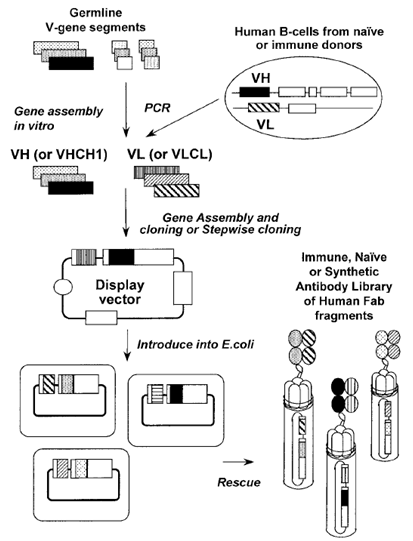
For library construction, large repertoires of single-chain Fv antibody fragment (scFv) or Fab antibody genes are cloned into engineered phage or phagemid vectors in a way that these antibody fragments can be displayed on the bacteriophage surface. One of the most successful applications of phage display has been the isolation of monoclonal antibodies from large random combinatorial phage antibody libraries (Fig. 1). Such libraries have been built in scFv and fab format by researchers. Typically, the antibody fragments are displayed on the surface of phage as either fab fragments, single-chain variable region fragments (scFvs), or dimeric scFvs, also known as diabodies, which differ from scFvs in the reduced length of the linker peptide used and their preference to associate as dimers.
Figure 1 Construction of a human Ab library displayed on phage. cDNA encoding for the heavy and the light variable regions of Abs (VH, VL) are amplified from human B cells by PCR and assembled. The assembled genes are inserted into a phagemid vector in frame with the gene encoding the CP pIII. The vector is introduced into E. coli. After rescue with helper phage, the random combinatorial library of Abs is displayed on phage and selection can be performed(Hennie R. Hoogenboom.etc).
The standard procedure of constructing antibody library is that:
1. Construction of Phage-Display Vector (choose suitable vector based on antibody format ).
2. Preparation of the cDNA Template: After the isolation of mRNA from the desired cell type and the preparation of cDNA, the construction of immune libraries is usually done by a two-step cloning or assembly PCR (choose appropriate clone methods based on library type).
3. Amplification of Antibody Variable Region Genes (choose proper PCR strategy and optimizing PCR reaction based on targeting sequence size and primer design requirement).
4. Construction of the scFv/fab Library (after target genes (scfv/fab genes) and optimized vectors are digested by same restriction enzymes, ligate their purified segments).
5. Rescue of scFv-Phage.
6. Panning of scFv/fab-Phage: The library was subjected to three or four rounds of panning. Notably, more than 90% of the selected clones showed positive binding for their respective antigens after as few as three rounds of panning.
7. ELISA and purification of scFv/fab-Phage Binding: choose proper antigen and expression system is the key factor of success.
To gain a high-quality library, firstly, the individual steps were optimized to increase the antibody variable region gene diversity and the efficiency of scFv/fab gene assembly and cloning. The primer design was optimized for amplification of variable region gene pools to maintain maximum diversity. Second, about scFv, the efficiency of scFv gene assembly was increased by exploiting the presence of the DNA encoding the (G4S)3 peptide linker included at the end of the 3’-primer of the VH gene and 5’-primer of the VL gene. This design allowed us to assemble scFv genes from only two DNA fragments. Several parameters about human universal libraries are given in Table 1.
Table 1 Human “single-pot”/universal phage display libraries.
| Library vector(library name) | Library type | Antibody type | Library cloning strategy | Library size |
| fdDOG-2lox,pUC19-2lox | Semisynthetic | Fab | PCR with random CDR3 primer, Cre-lox | 6.5 x 1010 |
| fdTet | Naïve | scFv | Recloning of a naïve library | 5 x 108 |
| pComb3 | Semisynthetic antitetanus Ab framework | Fab | PCR with random CDR H3 primer | >108 |
| pMorph series (HuCAL library) | Synthetic | scFv | PCR with random CDR H3 primer | 2 x 109 |
| pR2 (DAb library) | Semisynthetic | VH dAb | Assembly PCR random CDR VH | 3 x 109 |
| pMorph series (HuCAL GOLD library) | Synthetic | Fab | Two-step cloning, all CDR replacement | 1.7 x 107 |
| pSEX81 | Naïve | cFvs (with N-terminus of CH1 and CL) | Two-step cloning | 4 x 109 |
In summary, antibodies with subnanomolar affinities can be selected from either type of library, naïve or synthetic. If the assembly by cloning or PCR and preservation of molecular complexity is carefully controlled at every step of its construction, high-quality libraries of more than 1010 independent clones can be generated, with the potential to yield hundreds of different antibody fragments to any given antigen. The library yielded antibodies with in some cases nanomolar affinity against numerous different antigens. ScFv phage library proved to be difficult to re-propagate without significant loss of diversity. However, a more stable, large (1.2 x 109) clone scFv phagemid library made by standard cloning methods and using the synthetic V-genes, was recently shown to be equally effective.
Methodologies for the creation and selection of combinatorial antibody libraries on the surface of phage continue to be improved and extended. Now new library methodologies offer many alternative and at least as powerful routes, by combining the generation of billions of components with a fast screening or selection procedure to identify the most interesting lead candidates. Antibodies have been selected to molecules both large and small to study protein recognition, immunity, and problems in basic and applied sciences. In conclusion, antibody phage display will continue to be a main method for the generation of human therapeutic antibodies in the near future.