TCR is expressed on the surface of T cell and recognizes diverse antigens such as proteins, polysaccharides, bacteria or viruses. Usually, the TCR gene includes an α-chain and a β-chain, the former consists of variable (V), joining (J) and constant (C) gene fragments, while the TCR β-chain is made up of a V, J and C region, with an additional diversity (D) region and these genes can be grouped into a distinct TCR Vβ gene family. What's more, the rearrangement of those TCR genes plays a crucial role in antigen recognition by T cells, which enables T cells to recognize antigens specifically, and any dysregulation in this complex yet highly regulated process may lead to disease. Therefore, in T-cell immunotherapy development, the analysis of TCR repertoire clonality and diversity can serve as an important approach for evaluating virus, tumor or self-immune antigens-induced T lymphocytes clone types. Moreover, current evidence demonstrates that fast off-rates and serial engagement of MHC-peptide and TCR complexes may be crucial for T-cell activation, and tetramerized MHC class I and class II complexes have been used extensively to quantify and characterize antigen-specific T cells and to probe TCR-MHC interaction. The accessibility of various MHC-peptide tetramers can be great helpful for the study of specific TCR.
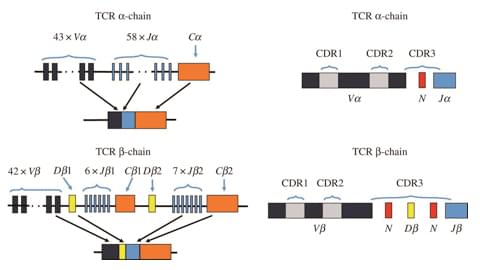
As a leading custom service provider in the field of T cell receptor (TCR) development, Creative Biolabs has highly qualified groups, advanced technologies and wide platforms to perform the most comprehensive TCR analysis, and our first-class TCR assay system covers the assessment of TCR clonality, the analysis of CDR3 diversity as well as the TCR Vα/β repertoire evaluation. We are committed to providing our customers accurate assay data and help every customer develop better TCR products in a time-saving and cost effective way.
With full understanding of the function and working mechanism of TCR, scientists of Creative Biolabs carry out custom TCR analysis services which include TCR-repertoire sequencing, TCR Vβ Repertoire Analysis, MHC-peptide tetramer, TCR clonality assessment and Clonality quantitation and CDR3 size diversity determination. Meanwhile, our scientists have developed a novel assay technique to overcome the shortcoming of traditional techniques (e.g. flow cytometry and PCR) in TCR clonality assessment. We guarantee that reliable, rapid, sensitive and quantitative analysis will be performed throughout your whole TCR assay project. For each TCR analysis project, Creative Biolabs will present and deliver a comprehensive report including detailed procedures and results for your project, such as analytical methods, all analysis results, appropriate graphs and calculations. All raw data is also available upon request.
Based on our combinatory and comprehensive package of analytical methods, Creative Biolabs has come to the unique position to not only have the technology and experience necessary for all kinds of TCR analysis, but also the capability to provide these analytical services in the most cost and time efficient manner.
To discuss your demands or to request a proposal, please contact us by .
Use the resources in our library to help you understand your options and make critical decisions for your study.
TCR-T cell therapy is an emerging form of cancer immunotherapy that modifies T cells to recognize and attack cancer cells more effectively. T cells are genetically engineered to express high-affinity T-cell receptors (TCRs) that can bind to specific antigens on cancer cells, triggering immune responses to eliminate the tumors. While promising for treating myeloma, myeloid malignancies, and solid tumors, this therapy faces challenges including safety concerns, potential off-target effects, and the need for improved manufacturing processes. Ongoing clinical trials continue to refine TCR-T therapy, highlighting both its potential and the hurdles that must be overcome for widespread clinical application.
CAR-T cell therapies revolutionize cancer treatment by harnessing the power of a patient's immune cells. Engineered with chimeric antigen receptors (CARs), these cells effectively target and destroy cancer cells, offering a personalized and potent approach. This groundbreaking immunotherapy has shown remarkable success in treating certain blood cancers, providing hope for patients who may not respond to traditional treatments. CAR-T cell therapies mark a significant stride towards precision medicine, ushering in a new era in oncology with the potential to transform the landscape of cancer care.
Based on the significant roles of T cells in the immune system, many small-molecule drugs targeting T cells and T-cell based immunotherapies have been developed for the treatment of intractable diseases including autoimmune diseases and cancer. T cell-based immunotherapies mainly utilize the mechanisms of T cell-mediated immune responses and the effects of some other immune cells such as dendritic cells (DCs), natural killer (NK) cells, and macrophages.
Adoptive cell transfer (ACT) of engineered T cells is a cutting-edge therapeutic approach revolutionizing cancer treatment. This innovative method involves modifying T cells, a key component of the immune system, to enhance their ability to target and eliminate cancer cells. By introducing genetically engineered T cells into patients, researchers aim to bolster the immune response against cancer, offering a personalized and potentially curative treatment option. This groundbreaking technology holds promise for addressing various malignancies and represents a significant stride towards more effective and precise cancer therapies.
Reference
For any technical issues or product/service related questions, please leave your information below. Our team will contact you soon.
All products and services are For Research Use Only and CANNOT be used in the treatment or diagnosis of disease.
 NEWSLETTER
NEWSLETTER
The latest newsletter to introduce the latest breaking information, our site updates, field and other scientific news, important events, and insights from industry leaders
LEARN MORE NEWSLETTER NEW SOLUTION
NEW SOLUTION
CellRapeutics™ In Vivo Cell Engineering: One-stop in vivo T/B/NK cell and macrophage engineering services covering vectors construction to function verification.
LEARN MORE SOLUTION NOVEL TECHNOLOGY
NOVEL TECHNOLOGY
Silence™ CAR-T Cell: A novel platform to enhance CAR-T cell immunotherapy by combining RNAi technology to suppress genes that may impede CAR functionality.
LEARN MORE NOVEL TECHNOLOGY NEW SOLUTION
NEW SOLUTION
Canine CAR-T Therapy Development: From early target discovery, CAR design and construction, cell culture, and transfection, to in vitro and in vivo function validation.
LEARN MORE SOLUTION

