B Cell-based Small-angle Scattering Assay
Brief Summary
Small-angle X-ray scattering (SAXS) is a valuable structure analysis technique applied to acquire data on the shape, structure changes, and size of macromolecules in solution. SAXS is typically used to study the interactions between macromolecules and the functional mechanisms of molecules in cells. This method can offer low-resolution structural graphs of bound proteins, such as antigen-antibody complex, on the shape, conformation, and binding state.
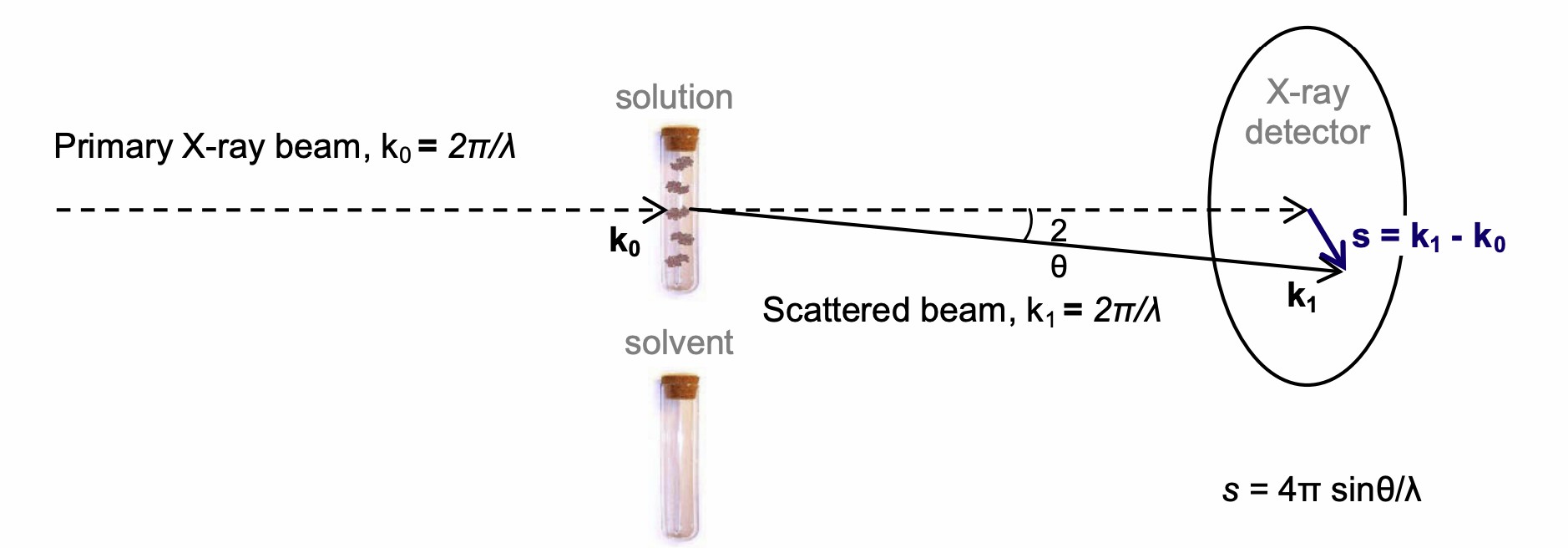 Fig.1 Schematic representation of a SAXS experiment.2
Fig.1 Schematic representation of a SAXS experiment.2
Cell-based Small-angle Scattering Assay Service at Creative Biolabs
The powerful X-ray scattering techniques provided us an insight into the complex interaction between the specific antibody and peptides. A SAS model of the antibody and peptides complex could indicate the shape information on the spatial arrangement of antibodies on an antigen, antibody binding sites, and conformation.
Workflow
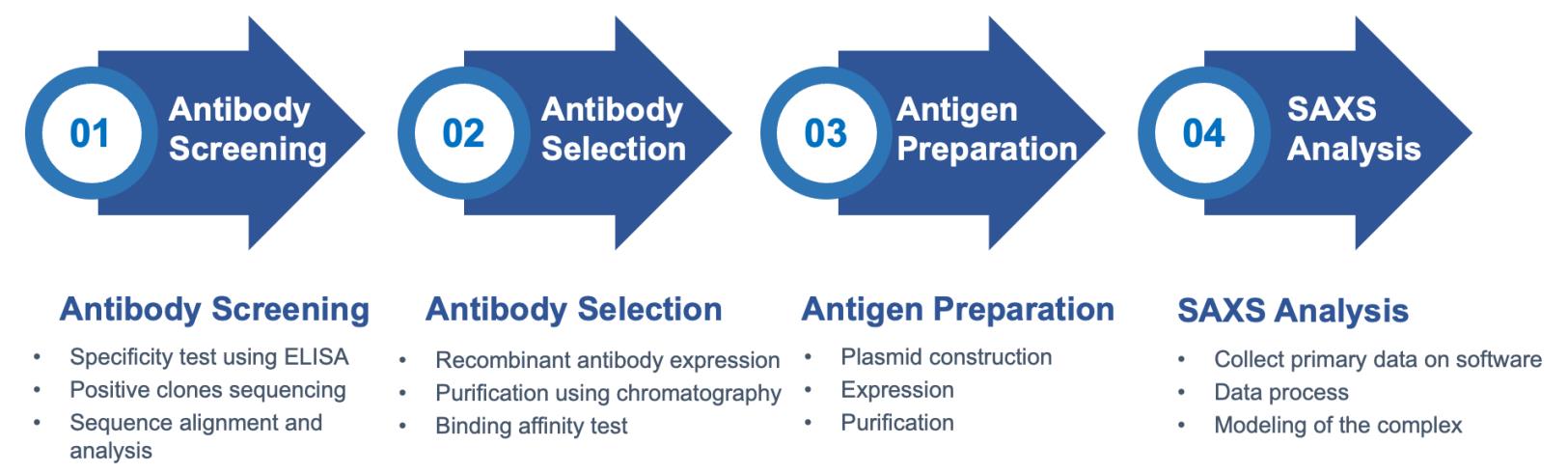
Key Features of the Small-angle Scattering Assay

Highlights

Our Capabilities
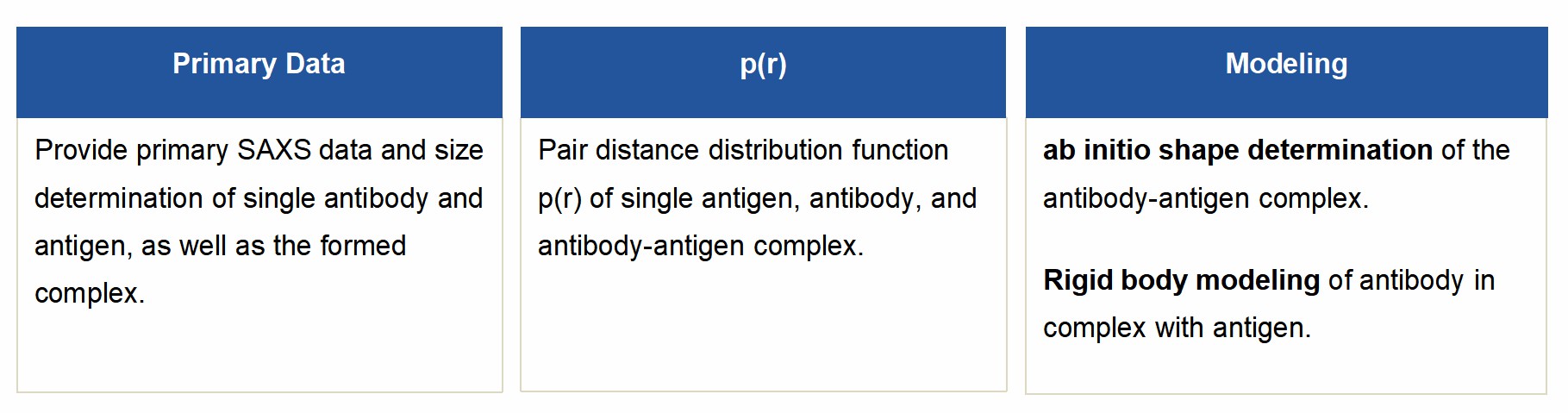

Data Analysis Software on Our Instrument
|
Function
|
Software
|
Description
|
|
Experimental data processing
|
PRIMUS, GNOM, CHROMIXS
|
Primary data processing
|
|
Ab initio modeling
|
MONSA, GASBOR, DAMMMIN, DAMMIF
|
Determine shape using dummy atom model
|
|
Rigid body modeling
|
CORAL, SASREF, BUNCH, GLOBSYMM
|
Model the multiple motif ensembled complexes
|
|
Structure refines
|
SREFLEX
|
Refine high-resolution protein models
|
|
Model evaluation and manipulation
|
CRYSOL, SRYSON, CIFSUP, DAMAVER, SASpy
|
Computationally model the scattering pattern
|
Case Study
|
Analysis of SARS-CoV-2 Novel Neutral Antibody Binding Site on Spike RBD
|
-
Background
Sb23 is a neutral antibody selected from the antibody library and binds to the RBD of spike protein on SARS-CoV-2.
|
-
Research Objectives
Structural basis of the neutralization activity of the antibody-Sb23 against SARS-CoV-2 RBD.
|
-
SAXS Assay Parameters
Instrument: EMBL P12 beamline of the storage ring PETRA III
Scatter detector: Pilatus-6M pixel
Protein solution concentration range: 0.3–4 mg/ml
|
-
Results
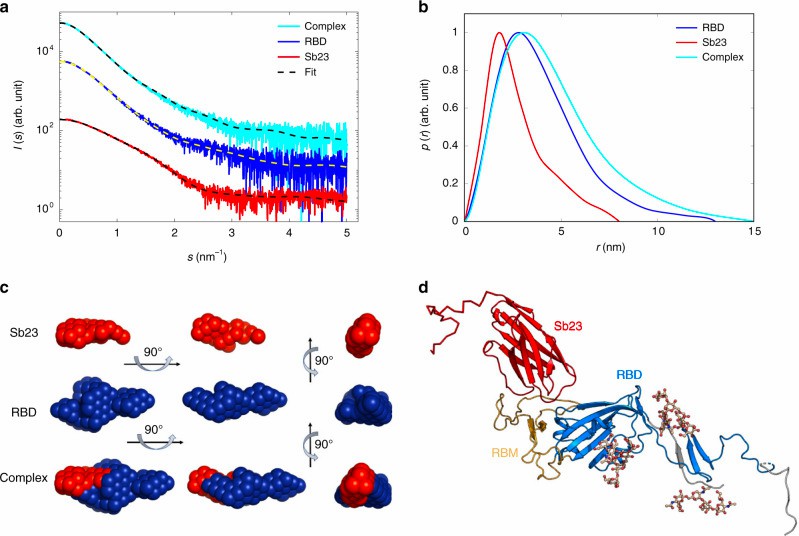
Fig.2 Applying SAXS to characterize the interaction between antibody-Sb23 and spike RBD.1
|
Creative Biolabs is pleased to support your cancer epitope analysis project. Please contact us and discuss your project with our highly skilled experts.
References
-
Custódio, T. F.; et al. Selection, biophysical and structural analysis of synthetic nanobodies that effectively neutralize SARS-CoV-2. Nature communications. 2020, 11(1): 5588.
-
Kikhney, A. G.; Svergun, D. I. A practical guide to small angle X-ray scattering (SAXS) of flexible and intrinsically disordered proteins. FEBS letters. 2015, 589(19 Pt A): 2570–2577.
-
Manalastas-Cantos, K. et al. ATSAS 3.0: expanded functionality and new tools for small-angle scattering data analysis. J. Appl. Cryst. 2021, 54: 343-355.
For Research Use Only | Not For Clinical Use


 Fig.1 Schematic representation of a SAXS experiment.2
Fig.1 Schematic representation of a SAXS experiment.2





 Download our brochure
Download our brochure

