Polymerase Chain Reaction (PCR)-based Tumor Profiling
Background
Tumor profiling involves the use of various technologies to understand and uncover the characteristics of cancer cells. Various technologies and platforms to profile genomic/transcriptomic/proteomic aberrations of cancers are aiming to guide precision treatment. The traditional molecular profiling method-polymerase chain reaction (PCR) allow for rapid, sensitive, and accurate detection of potential cancer biomarkers. The major drawback of the genome-wide application of PCR is low throughput. However, PCR is typically a good choice when the number of target regions is low and when the study aims are limited to screening or identification of known variants.
PCR-based Tumor Profiling Services
Polymerase chain reaction (PCR) has given researchers the ability to detect and identify mutations and gene expression in a variety of sample sources. Creative Biolabs provides reliable tumor gene expression profiling services to help you accelerate your research. Our PCR-based tumor profiling service is a rapid, cost-efficiency method for screening or identification of known gene expression.
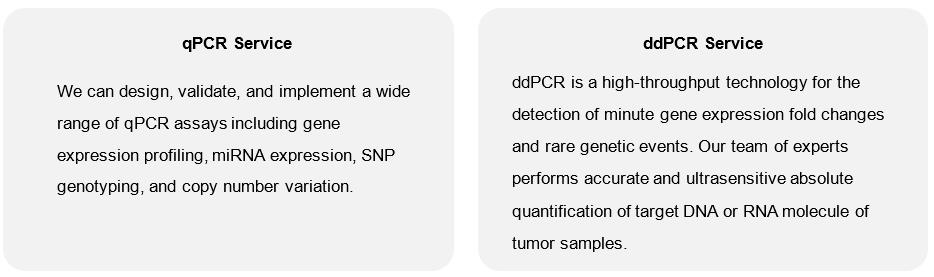
With over years of experience in integrated bioanalysis and assay development services, Creative Biolabs offers a well-established platform for rapid and reliable quantitative PCR analysis. Our team can support any quantitative PCR-based (qPCR) or droplet digital PCR (ddPCR) using off-the-shelf assays or validated custom assays. Our scientists will help you settle on a PCR services workflow that fits your needs and budget. The full-service option includes RNA/DNA extraction, reverse transcription, primer/probe design, qPCR/ddPCR reaction optimization, and complete data analysis.

Workflow
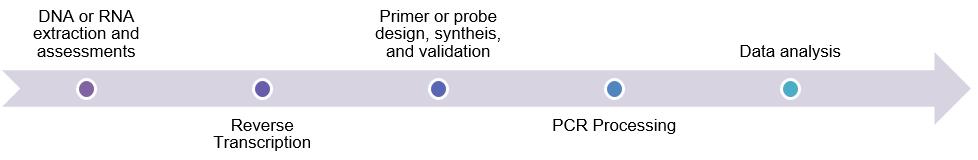

One-Stop Tumor Profiling Platform
As a leading biotechnology company, Creative Biolabs has extensive experience in multiple tumor profiling technologies including IHC, FISH/CISH, PCR, and Next-Generation Sequencing(ngs). We are dedicated to providing you with the most comprehensive and professional PCR services. Using proprietary tools, our team can custom-design PCR primers and probes to further support your experiment. Each client receives a complete study report including detailed experimental procedures, primer/probe sequences, raw data, and data analysis file with gene expression fold-change information. For more details, please feel free to contact us and we will discuss your requirements.
Frequently Asked Questions
Q1: Do I need to supply a positive and negative control?
A1: It is recommended to run a positive control and a negative template control such as RT reagent control without an RNA template added.
Q2: What sample types can be analyzed using qPCR?
A2: qPCR can be used to analyze many different biological sample types, including cultured cells, blood samples, fresh tissues, fixed-formalin paraffin-embedded (FFPE) samples, etc.
Q: What format will the data be delivered in?
A: We provide the raw data. For further analysis, you can contact us to discuss.
Published Data
Data 1: Detection and quantification of circulating tumor DNA in plasma of head and neck cancer patients.
Sample type: blood plasma samples
Tumor: head and neck cancer
Technology: ddPCR
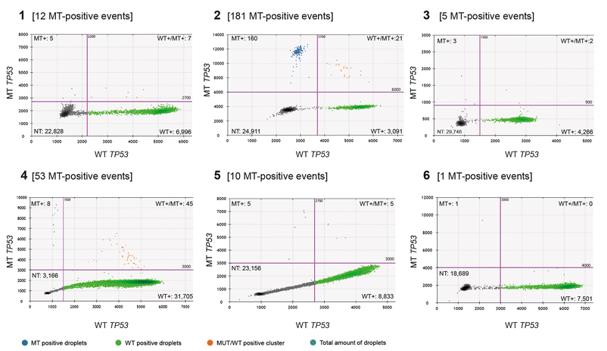 Fig.2 The result of TP53 mutations in ctDNA in plasma from patients using ddPCR.1
Fig.2 The result of TP53 mutations in ctDNA in plasma from patients using ddPCR.1
Contact Us
Have a PCR question? Our support team is on hand to help, get in touch!
Reference
-
van Ginkel, Joost H., et al. "Droplet digital PCR for detection and quantification of circulating tumor DNA in plasma of head and neck cancer patients." BMC cancer 17 (2017): 1-8. Distributed under Open Access license CC BY 4.0, without modification.
For Research Use Only | Not For Clinical Use




 Fig.2 The result of TP53 mutations in ctDNA in plasma from patients using ddPCR.1
Fig.2 The result of TP53 mutations in ctDNA in plasma from patients using ddPCR.1
 Download our brochure
Download our brochure