SuPrecision™ Sequencing Services

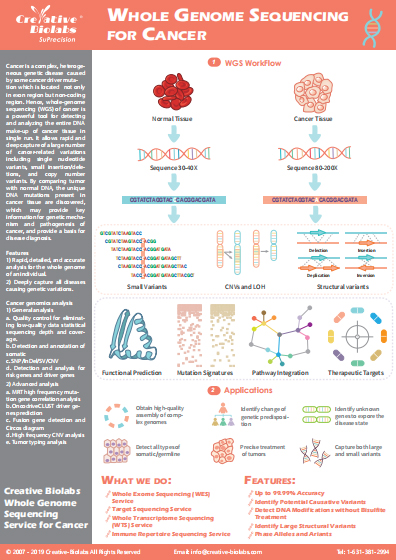
The development of WGS, combined with the falling cost of the technology holds great promise for accelerating the delivery of genetic predictors of human diseases and related treatment. WGS can be used to comprehensively explore all types of genomic alterations in cancer, to better understand the whole landscape of driver mutations and mutational signatures in cancer genomes and to elucidate the clinical implications of these unexplored genomic regions and mutational signatures. WGS is an important analysis platform for mutation interpretation and cancer precision medicine. Creative Biolabs has developed an efficient WGS Bioinformatics Analysis pipeline for WGS data analysis. We have accomplished many WGS projects for medical research in cancer and complex genetic disorders.

WES provides coverage of more than 95% of the exons, which harbor the majority of the genetic variants associated with human disease phenotypes. WES offers researchers the ability to use sequencing and analysis resources more efficiently by focusing on the most relevant portion of the genome and facilitates the discovery and validation of common and rare variants. WES is especially suitable for patients who are looking for a unifying diagnosis of multiple medical problems. Creative Biolabs has accumulated extensive experience in precision medicine WES program associated with corresponding WES Bioinformatic Analysis to study a cancer person’s exome and illustrate how this can be used to identify novel biomarkers associated with a response.
Targeted sequencing is an NGS technique focusing on amplicons and specific genes and it is an effective approach to diagnose various genetic disorders. Targeted sequencing technology has demonstrated its clinical utility as it provides the basis for precision medicine to be applied in the cancer patient. Creative Biolabs can provide a one-stop solution, from project consultation, experimental design to sequencing and comprehensive Targeted Sequencing Bioinformatics Analysis service.

WTS is an important tool for the understanding of the molecular mechanisms of many diseases including human cancers. Creative Biolabs offers a series of cancer WTS service, including but not limited to mRNA transcriptome sequencing (mRNA-Seq), microRNA sequencing (miRNA-seq), circular RNA sequencing (circRNA-seq) and long non-coding RNA sequencing (lncRNA-seq). Our scientists have accumulated extensive experience in providing the most accurate and comprehensive sequencing service and corresponding WTS Bioinformatic Analysis service.
The recent advent of high-throughput immune repertoire sequencing gives us direct insight into the diversity of B-cell and T-cell receptor (BCR and TCR) repertoires. Our scientists have accumulated extensive experience in identifying TCR (α, β, γ, and δ) and BCR V(D)J sequences contained in a single sample. Combining with our Immune Repertoire Bioinformatic Analysis pipeline, many data that indicate the great potential to change the way we diagnose, treat, and prevent immune system-related disorders, have been offered to our global customers. Accumulation of such knowledge will lead to the development of effective ways for personalized immune modulation through deeper understanding of the mechanisms that affect our immune ecosystem.

Epigenomics is a series of epigenetic modifications that affect gene activity and expression without altering the DNA sequence. Various epigenetic modifications, such as DNA methylation and histone modification, have been reported to be associated with the pathological process of cancer. We provide a a full spectrum of high-quality epigenomic sequencing and analysis services, thus accelerating a novel sight into the tumorigenesis.
The 3D configuration of the genome is crucial for gene regulation. It is complex and dynamic, and can vary between cell types and during cell differentiation and development. To understand the workings of the genome, we must study the chromosome structure. We use high-throughput chromosome conformation capture (Hi-C) technology to help analysis the role of chromatin 3D organization in gene expression.

Thousands of microbial species comprise the normal human microbiome. The microbe communities can substantially influence normal physiology and response to diseases, including cancer. Metagenomics studies in tumor samples have been paid more and more attention in pharmaceutical and biotechnological industries for the discovery of novel drugs. Metagenomics sequencing provides a powerful toll to broaden the scope of targeting microbes responsible for various types of cancers.
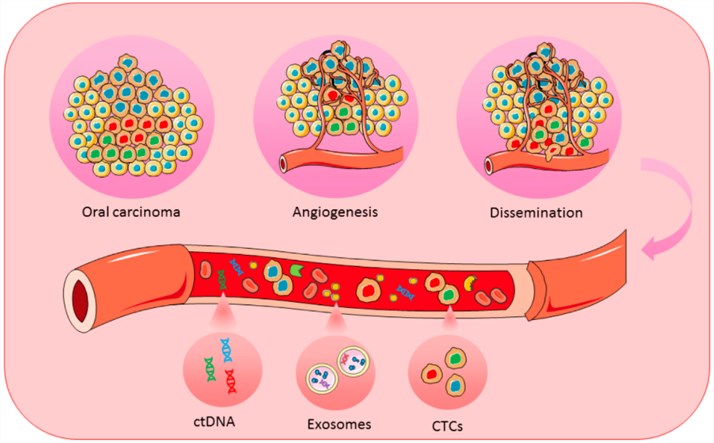
ctDNA testing, also known as liquid biopsy, is a less-invasive diagnostic alternative to tissue biopsies and is expected to be applied in tumor diagnosis and prognosis. NGS provides an excellent technology that allows the detection of low levels of ctDNA in the blood in a highly sensitive and specific manner. Creative Biolabs is a world-leading service provider that is committed to offering a full range of NGS-based DNA/RNA sequencing services including ctDNA sequencing for tumor research.

Single-cell sequencing technologies provide revolutionary tools for analyzing a single cell's gene expression, DNA variation, epigenetic state, and nuclear structure, thus improving the understanding of the molecular mechanisms for carcinogenesis and tumor progression.
 Distributed under CC BY-SA 4.0, from Wiki,
without modification.
Distributed under CC BY-SA 4.0, from Wiki,
without modification.
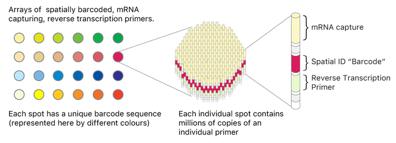
Spatial transcriptome sequencing technology provides a powerful tool for investigating the gene expression of individual cells in their original location and has been used in many research fields including pathology, immunology, developmental biology, neuroscience, and oncology.
Reference
- Lousada-Fernandez, Fatima, et al. "Liquid biopsy in oral cancer." International journal of molecular sciences 19.6 (2018): 1704. Distributed under open access license CC BY 4.0, without modification.