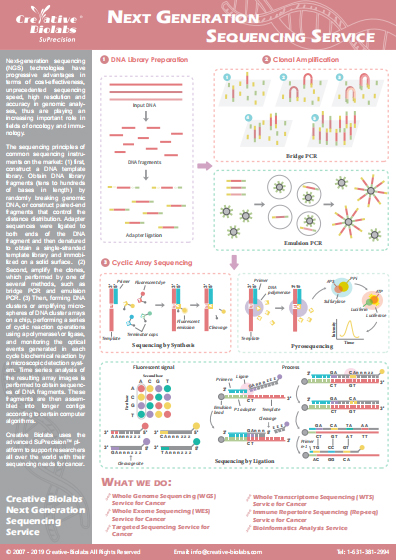
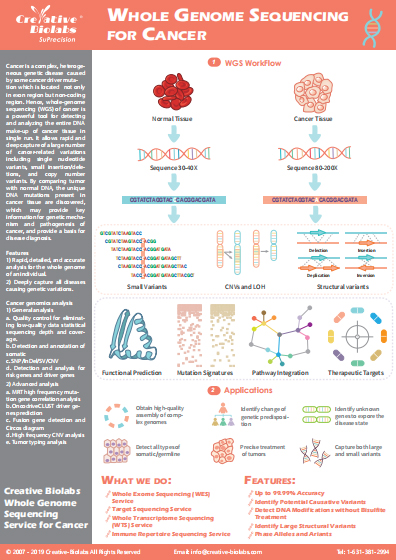
Small RNA Sequencing Service
Equipped with world-leading technology platforms and professional scientific staff, Creative Biolabs is a world-leading services provider in the field of next-generation sequencing (NGS). Based on our versatile SuPrecision™ platform, we offer one-stop small RNA sequencing services for global customers from small RNA extraction to data analysis. We are committed to delivering accurate, affordable and high-quality sequencing data to support your research.
Importance of Small RNA Sequencing in Cancer Research
Small RNA species are a class of lowly abundant, non-coding RNA molecules with a length of less than 200 nt. The kinds of small RNA include microRNA (miRNA), small interfering RNA (siRNA), Piwi-interacting RNA (piRNA), small nucleolar RNA (snoRNAs), tRNA-derived small RNA (tsRNA), small rDNA-derived RNA (srRNA) and small nuclear RNA (snRNA). These small RNAs play essential roles in gene silencing and post-transcriptional regulation of gene expression, involved in a number of biological processes including cell proliferation and differentiation, and apoptosis. Currently, accumulated evidence demonstrated that small RNAs are associated with the development and progression of cancer, as well as anti-tumor drug resistance. For example, the over-expression of miR-17 was associated with chemo-resistant colon cancer. miR-34 was involved in the regulation of drug resistance in gastric, prostate and breast cancers. The upregulated levels of piR-4987, piR-20365, piR-20485 and piR-20582 were found in tumor tissue. Hence, to explore small RNAs’ roles in oncology will increase the understanding of tumor pathological progression and accelerate discovery of novel anti-tumor therapies.
Small RNA-Seq is a powerful method that can quickly identify the known small RNAs’ level under specific conditions and discover novel small RNA molecules. Meanwhile, it allows small RNAs’ profiling under different conditions and the correlation analysis with transcriptome sequencing expression profile data.
Small RNA-Seq Workflow at Creative Biolabs
-
 Total RNA
Total RNA
Isolation
-
 Small RNA
Small RNA
Enrichment
-
 Small RNA Library Construction
Small RNA Library Construction -
 Sequencing
Sequencing
-
 Bioinformatics
Bioinformatics
Analysis
Sample Requirements
- Tissue or cell Sample:
1) Fresh cultured cell (number of cells): ≥ 2×105 cells
2) Fresh animal tissue (dry weight): ≥ 30 mg
3) Whole blood: ≥ 3 ml
4) FFPE: 5 pieces, 100 mm2, 5~10 μm thick, unstained
- RNA Sample:
1) Amounts of total RNA: ≥ 10 μg;
2) Concentration of total RNA: ≥ 200 ng/μL;
3) RNA purity: OD260/280=1.8~2.2, OD260/230 ≥ 2.0, RIN ≥ 6.5, 28S:18S ≥ 1.0;
4) Amounts of small RNA: ≥ 1 μg;
5) Concentration of small RNA: ≥ 20 ng/μL.
Note! There are differences in sample requirements between different samples. For details, please consult our scientists.
Bioinformatics Analysis

Standard Analyses
1) QC for raw data
Sequence alignment
2) Small RNA classification and annotation
3) Analysis for known miRNA (including expression levels, length distribution, and base preference)
4) Functional element annotation
5) Prediction for novel small RNA
6) Analysis of small RNA expression difference between samples
7) Target gene prediction
8) GO annotation and KEGG passway analysis for differentially expressed small RNA target gene
...
Senior Analyses
1) Prediction for miRNA target gene (non-model species or new miRNA)
2) Analysis of the differential expression of novel miRNA
3) Correlation analysis between differentially expressed miRNA and differentially expressed target genes
4) Interaction network between differentially expressed miRNA and differentially expressed target genes
5) Functional clustering analysis of interaction target genes (KEGG and GO)
...
Advantages
- More than 3 million sequences can be obtained by one sequencing
- Not only identify the known small RNA but also discover novel small RNA
- Enable to detect the difference of single base of small RNA
- High reliable and repeatable data
Creative Biolabs provides small RNA sequencing service packages including sample preparation, library construction, deep sequencing, raw data quality control, and customized bioinformatics analysis. If you have additional requirements or questions, please feel free to contact us.
Q&As
Resources
Infographics
Podcast
- mRNA Sequencing
- CircRNA Sequencing
- LncRNA Sequencing
- Exosomal RNA Sequencing
- Ultra Low Input RNA Sequencing
- FFPE Tissue RNA Sequencing