Creative Biolabs offers academic and industrial researches on the high resolution paratope mapping of antibody all over the world, depending on novel research methods and tools. We are capable of meeting the specific need of each customer with the best project design.
Paratope mapping is characterization of the binding sites, or paratopes of antibodies which recognized by antigens. However, to develop vaccines and bio-therapeutics, one of the most important processes is identification of the antibody paratopes. There are a number of crucial uses to understand the paratope in the antibody, for example, predicting suitable antigens as vaccine components, improving our knowledge of the immune response and autoimmunity, gaining an supernormal understanding of a therapeutic antibody’s mechanism of action.
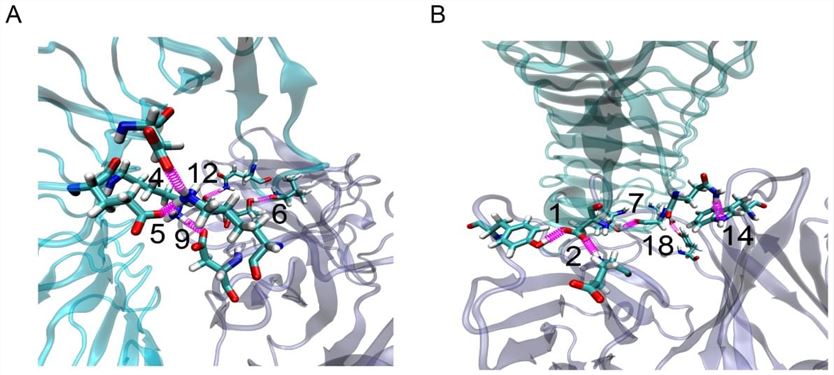 Fig.1 Paratope Mapping using computational procedure (Xiang Fang et al.,2012).
Fig.1 Paratope Mapping using computational procedure (Xiang Fang et al.,2012).
Compared with conventional approaches, Creative Biolabs has developed a number of novel research methods to perform paratope mapping, which is lower cost and faster, including:
Rigid body docking, free and steered molecular dynamics (MD) simulations and homology modeling constitute the computational procedure. With the computational procedure, it is reachable to identify key paratope residues on the target antibody and their partners on antigen. The computational model of quaternary structure of complexes consisted of more than two biological macromolecules which interact with each other is macromolecular docking. Molecular dynamics (MD) is an analysis to research the motion of molecules and atoms. To overview the dynamical evolution of this system, molecules and atoms are performed to interact for a certain time. Comparative modeling of protein, termed homology modeling as well, builds an atomic-resolution models of the protein from its amino acid sequence, and furthermore builds an experimental three-dimensional structure of a correlative isogenous protein.
By using Hydrogen-deuterium exchange and mass spectrometry, in the solvent hydrogen atoms will exchanged by backbone amide hydrogen atoms and if there is deuterium in the solvent deuterium but don’t contains hydrogen then the same exchange process occurs so that the protein can be deuterated. The flexibility and the availability of the target part of the protein in the solvent decide the rate of any special hydrogen atom happens. The hydrophobic core of a rigid protein where hid hydrogens will exchange at all, while those on a flexible loop will exchange nearly instantaneously.
Peptide-based approaches are based on the composition of overlap peptides, however the peptides must contain the entire sequence of the antigen. Following the peptides fixated onto a solid surface as an array and the binding to the antibody of curious is determined in an ELISA format. This approach can make antibodies ‘panned’ with very huge phage libraries available. Furthermore, the recombinant peptide genes from binding phage particles can be sequenced to identify the sequences and putative epitopes.
Using a Hopfield Network while representing the epitope and paratope in a complex as 3-dimensional attributed graphs, a mapping is estimated from a random training sample while representing. With the help of KBCM when using Hopfield Network, mapping paratopes on immunoglobulins to the epitopes can be available.
Creative Biolabs provides other various Epitope mapping services. Please feel free to contact usfor a detailed quote.
References
All listed services and products are For Research Use Only. Do Not use in any diagnostic or therapeutic applications.