Creative Biolabs now provides first-class membrane protein production service using E. coli cells system. Equipped with leading technical platforms and experienced scientists, we guarantee our clients the most satisfactory membrane protein products to deliver various research goals.
E. coli (Escherichia coli) is the most widely used organisms for the production of recombinant proteins. Its well-established application as a cell factory started since sufficient knowledge has been acquired about its physiology, genetics and genome sequence. Until now, E. coli expression system is still one of the most important heterologous systems to produce many different types of recombinant proteins.
Among E. coli expression systems, the T7 system is by far the most popular approach for producing proteins. It is reported that a derivative strain of E. coli, termed Lemo21 (DE3), is engineered in which the activity of the T7 RNA polymerase can be precisely controlled by its natural inhibitor T7 lysozyme (T7 Lys). In Lemo21 (DE3) strain, a large amount of recombinant membrane protein can be acquired easily. The development of this strain is sped up tremendously by using GFP-fusions to monitor membrane protein production. Strikingly, the Lemo21 (DE3) strain can be used to produce membrane protein GFP-fusions from which properly folded membrane protein will be recovered for further functional and structural studies (Figure 1).
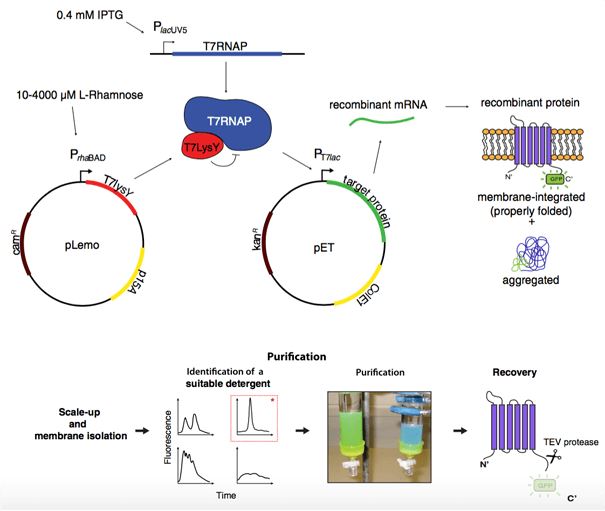 Figure 1. Genal procedures of membrane protein production yields in E. coli cells. (Biochim. Biophys. Acta., 2014)
Figure 1. Genal procedures of membrane protein production yields in E. coli cells. (Biochim. Biophys. Acta., 2014)
The advantages of using E. coli as the host organism are well known:
We have developed BacTEC™ and FoldEZ™ technologies to produce many hard-to-express and hard-to-dissolve proteins. In virtue of our BacTEC™ technology, the expression level can be significantly improved. In addition, over 98% of the inclusion bodies can be refolded and solubilized by our proprietary FoldEZ™ protein refolding technology. Besides E. coli, Creative Biolabs can select other bacterias as membrane protein production hosts, such as Lactococcus lactis (L. lactis), Bacillus subtilis (B. subtilis), and Rodhobacter sphaeroides (R. sphaeroides).
Creative Biolabs has been a long-term expert in producing a wide variety of membrane proteins based on our well-recognized E. coli expression system. Our profound expertise and strong beliefs to strive for excellence have earned us international reputation. We also provide multiple eukaryotic cell-based expression services as alternatives, including:
We look forward to assisting our global clients with diverse membrane protein projects through our capabilities and expertise. For more information, please feel free to inquire us.
References
All listed services and products are For Research Use Only. Do Not use in any diagnostic or therapeutic applications.