Single-nucleotide polymorphisms (SNPs) are DNA sequence variations usually caused by point mutations that give rise to different alleles containing alternative bases at a given nucleotide position within a locus. SNPs are considered the most useful biomarkers to confirm the inheritance of the parental wild-type or mutant allele thus increasing the accuracy of disease in vitro diagnostics (IVD) diagnosis and prognosis. Known for our deeply-rooted IVD expertise and abundant experience, Creative Biolabs is dedicated to offering a comprehensive range of SNPs analysis services to accelerate your IVD projects.
Brief Introduction of SNPs
SNPs are single genetic code variations that are the most common forms of genetic variation among people. Usually, SNPs are biallelic (two alternative bases occur) and have a minimum frequency (>1%) in the population. SNPs may contribute to changes in the genomic sequence, either in the coding (exons), intergenic, or non-coding (introns) region, therefore, they can influence mRNA splicing, nucleo-cytoplasmic export, stability, and translation. Due to their common frequency, ease of analysis, low genotyping costs, and the possibility to perform association studies based on statistical and bioinformatics tools, SNPs have gained increasing importance as biomarkers for disease IVD diagnosis and prognosis.
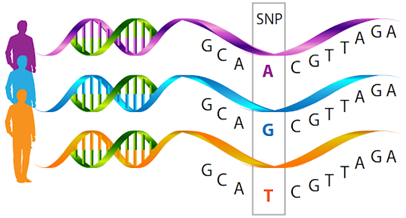 Fig.1 DNA microarray heatmap.Distributed under CC BY 4.0, from Wiki, without modification.
Fig.1 DNA microarray heatmap.Distributed under CC BY 4.0, from Wiki, without modification.
Methods for SNP Analysis
There are a handful of molecular strategies have been discovered for SNP analysis. These methods start with the common step -target sequence amplification, and subsequent 1) distinction of DNA sequence variants by short hybridization probes or by restriction endonucleases; 2) discrimination of mismatched DNA substrates by polymerases or ligases; or 3) by observing the template-dependent choice of nucleotide incorporated by a polymerase. Recently, assay formats that simplify scale-up in SNP analysis are also widely used.
Table.1 SNP analysis methods.
| Method | Allele discrimination by | Format | Instrumentation |
| Multiplexed allele-specific diagnostic assay (MASDA) | Allele-specific oligonucleotide (ASO) | format 1 array + gel | none |
| Whitehead-Affymetrix SNP microarray | ASO | format 2 array | array scanner |
| RT-PCR | ASO | homogeneous closed tube | RT-PCR instrument |
| Molecular beacons | ASO | homogeneous closed tube | RT-PCR instrument |
| MADGE | Restriction cleavage | gel | none |
| Allele-specific PCR | 3' matched/mismatched primer | gel | none |
| Ligation assay (OLA) | ligation | solid-phase microtiter readout | fluorospectrophotometer |
| Solid-phase minisequencing | nucleotide incorporation | solid-phase microtiter readout | scintillation counter |
| Minisequencing on array | nucleotide incorporation | format 2 array | PhosphorImager |
| Minisequencing multiplex gel | nucleotide incorporation | solid phase + gel | automated sequencer |
| Minisequencing with FRET | nucleotide incorporation | homogeneous reagent addition | RT-PCR instrument |
| Pyrominisequencing | nucleotide incorporation | homogenous reagent addition | luminometer |
Format 1 and 2: ASO hybridizations are performed with either the DNA samples or the oligonucleotides arrayed on a surface, respectively.
SNPs Analysis Services at Creative Biolabs
SNPs analysis helps us understand the rules that govern the changes caused by genetic mutations which would pave the way for personalized medicine and disease in vitro diagnostics. With years of experience in SNPs analysis and genotyping, Creative Biolabs has developed and introduced a series of methods and technologies for SNPs analysis, such as PCR and Sanger sequencing-based solution, next-generation sequencing (NGS) and analysis, restriction fragment length polymorphism, random amplified polymorphic DNA analysis, hybridization assays, minisequencing analysis, etc. Our one-stop solution can also be customized to meet different levels of sensitivity, specificity and accuracy. You can choose our SNPs analysis services to accelerate your research in molecular diagnostic, clinical examination, drug design and individual medical diagnosis.

SNPs Analysis Case Study
SNPs serve as biological markers for pinpointing a disease on the human genome map and can be used to search for and isolate the disease-causing gene. Researchers reported a genome-wide homozygosity mapping study using high-density single nucleotide polymorphism (SNP) arrays to localize the Salla Disease locus to a 14.9 Mb region on chromosome 6. They performed SNP genotypes from two affected individuals and their parents to reveal the polymorphisms of SLC17A5, the gene encoding the lysosomal N-acetylneuraminic acid transport protein. Direct sequencing of SLC17A5 in the proband revealed homozygosity for the 115C-T (R39C) sequence variant, the common cause of Salla disease in Finland. Genome-wide SNP arrays with relatively high marker density showed great promising as a diagnostic tool.
SNPs analysis is explored to help in identifying susceptibility genes for complex genetic diseases (e.g., Alzheimer's Disease, Breast Cancer and Coronary Heart Disease), and making whole-genome association studies more feasible. SNPs analysis also contributed to the detection of drug susceptibility genes to help the pipeline of drug development. Based on our advanced technology and abundant experience, Creative Biolabs offers high-quality customized service covering the entire SNPs analysis process to best suit your technical, program, and budget requirements. If you want to get more information about our SNPs analysis, please feel free to contact us.
Published Data
1. Analysis of Gene SNPs in COVID-19 Disease Highlighting the Susceptibility and the Severity of the Infection
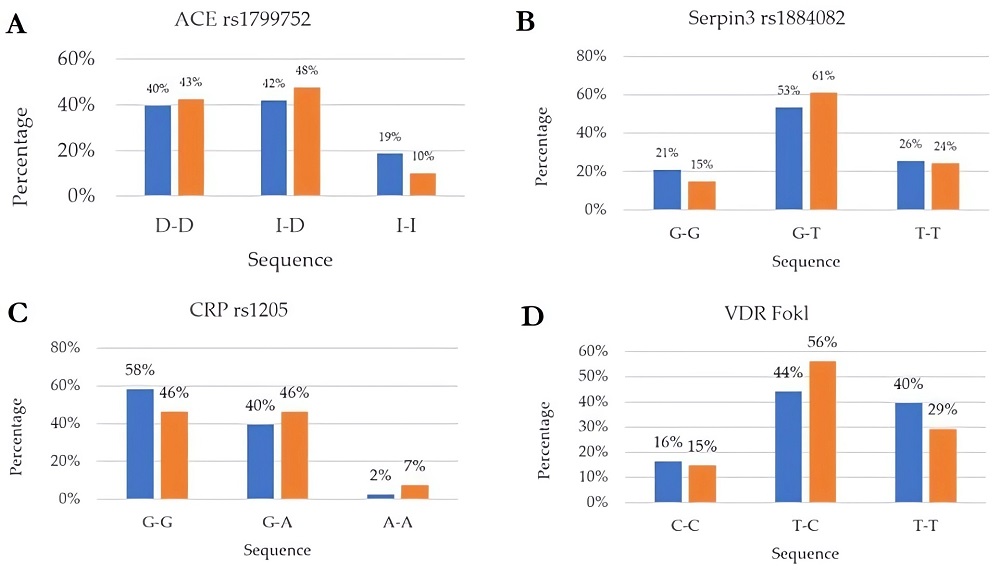 Fig.2 Genotype distribution of several SNPs among the healthy (blue) and affected (Orange) subjects analyzed.1
Fig.2 Genotype distribution of several SNPs among the healthy (blue) and affected (Orange) subjects analyzed.1
In this study, researchers investigated the presence of multiple genetic SNPs, the peripheral blood level of IL6, vitamin D, and arterial blood gas (ABG) markers in two distinct groups: COVID-19 patients (n = 41, study group) and healthy individuals (n = 43, control group). They analyzed genes related to pro-inflammatory responses and immune regulations, focusing on those implicated in the progression and complications of COVID-19. Key genes selected included IL1β, IL1RN, IL10, IL6, IL6R, TNFα, IFNγ, ACE2, SERPINA3, CRP and VDR (vitamin D receptor Tak1, Bsm1, and Fok1). Despite some limitations, the results suggested that various gene SNPs may be associated with a higher risk of COVID-19 infection. To conclude, understanding individual polymorphisms can help explain COVID-19 outcomes, promoting genetic profiling to develop personalized therapies and improve treatment strategies.
References
- Balzanelli, Mario Giosuè, et al. "Analysis of gene single nucleotide polymorphisms in COVID-19 disease highlighting the susceptibility and the severity towards the infection." Diagnostics 12.11 (2022): 2824. Distributed under Open Access license CC BY 4.0. The image was modified by extracting and using only part of the original image.
For Research Use Only.