Product List Background Lung Adenocarcinoma Aptamer Analysis
Background
Overview of Lung Adenocarcinoma
Lung adenocarcinoma, the most prevalent type of lung cancer, falls under the category of non-small cell lung cancer and is marked by distinct cellular and molecular characteristics. Patients typically present with persistent cough and shortness of breath, symptoms common to other lung cancer types. Originating in glandular cells that secrete mucus, lung adenocarcinoma often develops in smaller airways like alveoli and is usually located along the lung's periphery. This type of cancer generally exhibits slower growth compared to other lung cancers.
Lung Adenocarcinoma Related Signaling Pathways
Genetic alterations in lung adenocarcinoma often involve genes in the MAPK, p53, Wnt, cell cycle, and mTOR pathways. Over 80% of sequenced tumors have at least one mutation in the MAPK pathway, highlighting its crucial role in lung cancer. Mutations in PTEN, PI3K, and AKT genes, components of the insulin/PI3K/AKT signaling axis, are also prevalent. Additionally, dysregulation of mTOR is significant in lung carcinogenesis, presenting a potential therapeutic target. Further testing is needed to assess the efficacy of rapamycin and its analogues in treating lung adenocarcinoma.
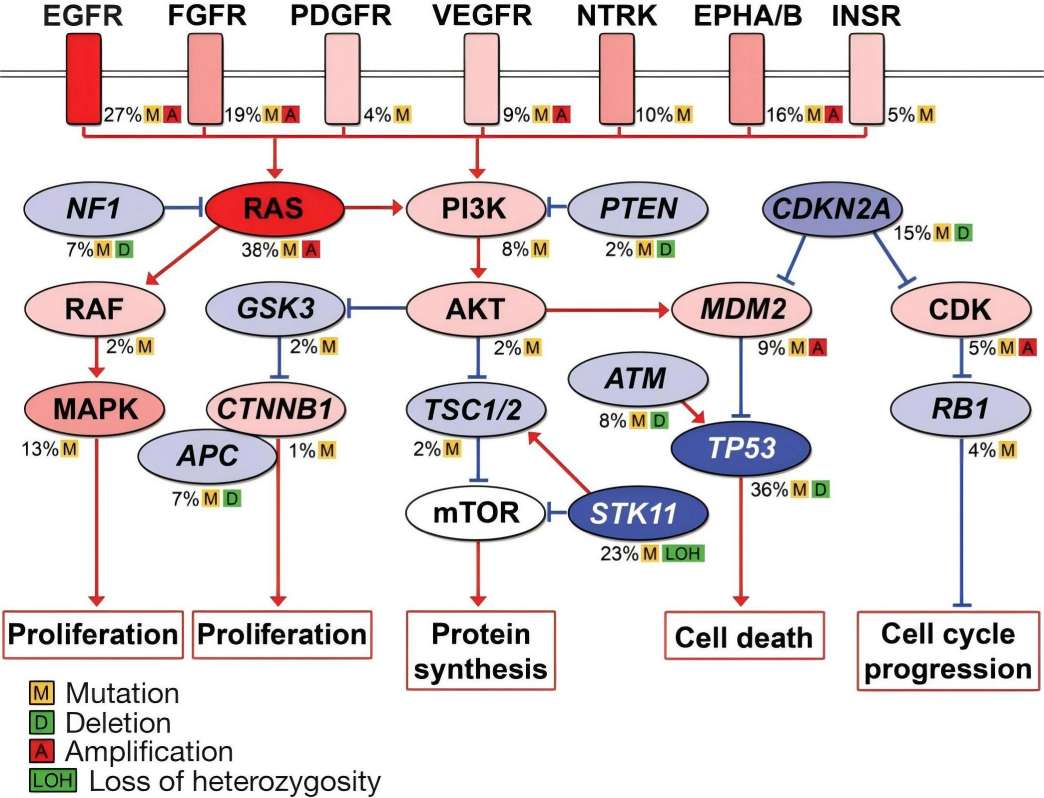 Fig. 1 Key mutated pathways in lung adenocarcinomas.1,4
Fig. 1 Key mutated pathways in lung adenocarcinomas.1,4
Utilizations of Lung Adenocarcinoma-Targeted Products
Aptamer-Based Detection of CTCs in Lung Adenocarcinoma
Circulating tumor cells (CTCs) serve as valuable clinical markers for metastasis prognosis and patient survival prediction. Traditional CTC analyses rely on antibody-based detection of epithelial markers, overlooking mesenchymal cancer cells in circulation. Researchers have selected and identified DNA aptamers as specific probes binding to lung adenocarcinoma cells from postoperative tissues, without binding to normal lung cells or lymphocytes. These aptamers are used to detect CTCs, apoptotic bodies, and microemboli in peripheral blood samples of lung cancer patients. Aptamer-associated protein biomarkers have been identified, enabling the production of tumor-specific aptamers for personalized diagnostics throughout anticancer therapy.
Anti-Lung Adenocarcinoma Tissue Aptamer as Molecular Probe and Therapeutic Method
Nucleic acid aptamers are increasingly utilized as molecular probes for pathology identification and imaging, and as platforms for targeted therapy. Recent studies demonstrate that aptamers can effectively characterize tumors, akin to monoclonal antibodies. Researchers have discovered DNA aptamers that bind effectively to altered lung cancer tissues, encompassing vascular networks, supportive connective tissues, and malignant cells. Protein targets are revealed using affinity purification and mass spectrometry, validated with antibodies and 3D imaging. These proteins are crucial for cancer progression, particularly lung adenocarcinoma. The aptamers can be used for aptahistochemistry, differential diagnostics, targeted therapy, intraoperative tumor visualization, and margin assessment.
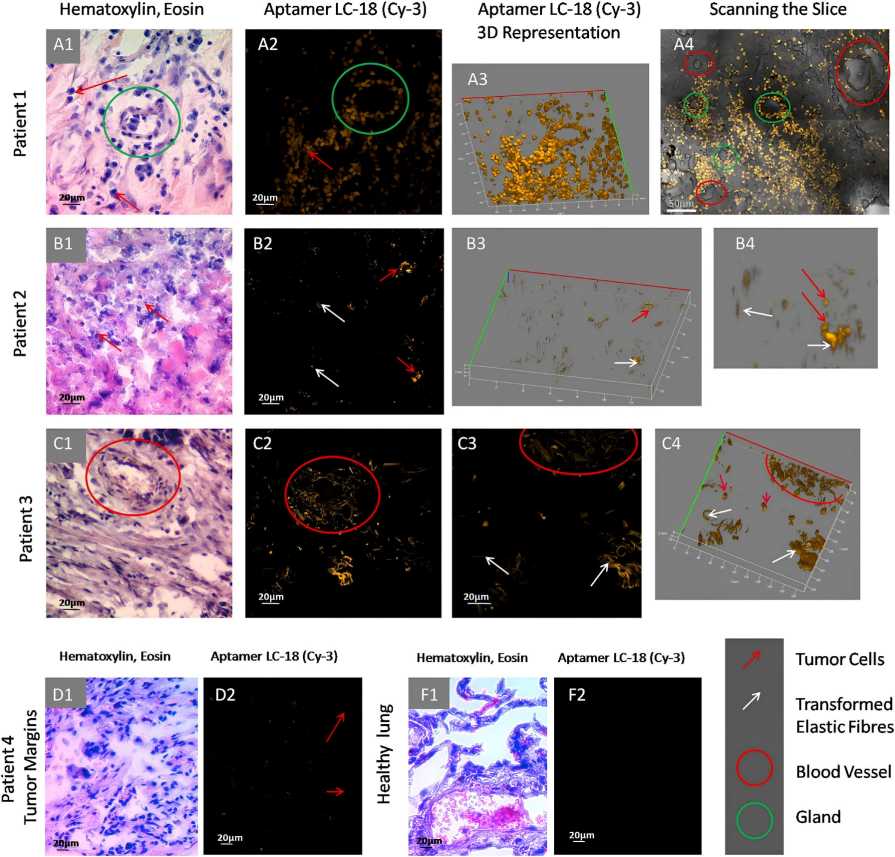 Fig.2 Microscopy imaging of anti-lung adenocarcinoma tissue aptamer binding to transformed elastic fibers in tumor blood vessels.2,4
Fig.2 Microscopy imaging of anti-lung adenocarcinoma tissue aptamer binding to transformed elastic fibers in tumor blood vessels.2,4
Lung Adenocarcinoma Aptamer Analysis
Creative Biolabs provides a wide range of lung adenocarcinoma-related products, including anti-lung adenocarcinoma aptamer. These products can effectively help to carry out your experiments and thus play an
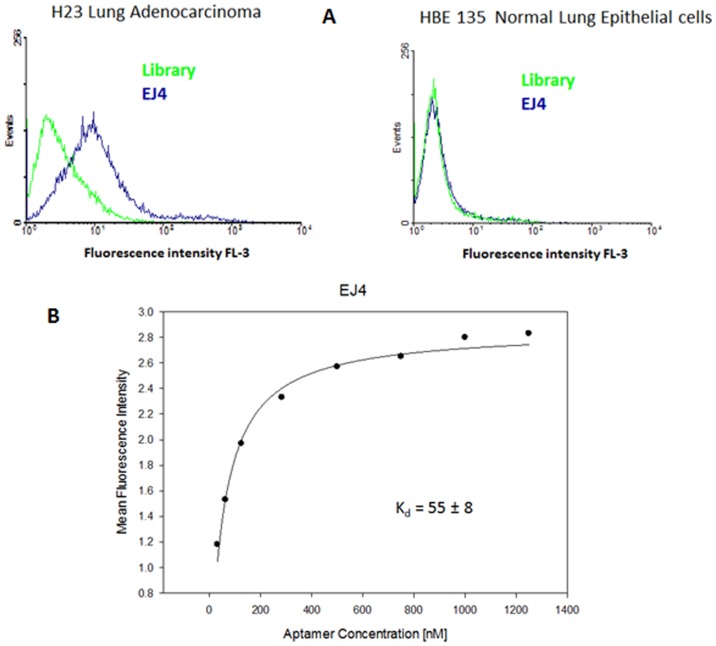 Fig. 3 Characterization of selected aptamers.3,4
Fig. 3 Characterization of selected aptamers.3,4
Elizabeth Jiménez et al. selected a set of aptamers that can distinguish lung adenocarcinoma cells from normal lung epithelial cells by the SELEX method.3,4 The study used H23 lung adenocarcinoma as a positive cell line, and HBE 135-E6/E7 normal lung epithelial cells as negative cell lines. The initial ssDNA random library that can recognize H23 cell surface membrane proteins was enriched by sequential binding to target cells, elution, and then 18 rounds of PCR amplification. In this process, in order to remove sequences that may bind to common proteins, reverse selection was introduced. When the selection was carried out to the 17th and 18th rounds, the fluorescence intensity observed by flow cytometry increased to a stable state, indicating that these aptamer candidate sequences can effectively bind to the H23 cell surface.
Creative Biolabs offers lung adenocarcinoma aptamer analysis, including SELEX service and other tailored functional services for our esteemed clients engaged in clinical and scientific research.
References
-
Ding, Li, et al. "Somatic mutations affect key pathways in lung adenocarcinoma." Nature 455.7216 (2008): 1069-1075.
-
Zamay, Galina S., et al. "DNA aptamers for the characterization of histological structure of lung adenocarcinoma." Molecular Therapy-Nucleic Acids 6 (2017): 150-162.
-
Jimenez, Elizabeth, et al. "Generation of lung adenocarcinoma DNA aptamers for cancer studies." (2012): e46222.
-
Distributed under Open Access license CC BY 4.0, without modification.


 Datasheet
Datasheet Fig. 1 Key mutated pathways in lung adenocarcinomas.1,4
Fig. 1 Key mutated pathways in lung adenocarcinomas.1,4
 Fig.2 Microscopy imaging of anti-lung adenocarcinoma tissue aptamer binding to transformed elastic fibers in tumor blood vessels.2,4
Fig.2 Microscopy imaging of anti-lung adenocarcinoma tissue aptamer binding to transformed elastic fibers in tumor blood vessels.2,4
 Fig. 3 Characterization of selected aptamers.3,4
Fig. 3 Characterization of selected aptamers.3,4
