Creative Biolabs offers customers a novel ribosome display method, the RIDS, for diverse studies in protein engineering.
Protein engineering is the process of designing and developing peptides or proteins with desired properties based on understood structure-function relationships. It mainly relies on the technologies in molecular biology, bioinformatics and biochemistry.
For the traditional ribosome display methods, the ribosome forms a stable complex with the translated protein and the mRNA that encodes the protein, and the release of the protein from the complex is slowed by the removal of a termination codon. Several display strategies have provided powerful and efficient techniques for the selection and evolution of peptides and proteins. In common protein display techniques, the coupling of genotype and phenotype is the single most critical determinant: the specific sequence information (genotype) of members of libraries that encode the selected protein (phenotype) can be determined from the corresponding DNA/RNA introduced into the system. The gene encoding the selected protein can then be re-amplified for further evolution and analysis. The methods have been used with success in many protein engineering studies, but they are associated with certain limitations and disadvantages as well. As cell-dependent display systems, they include a necessary in vivo step. Therefore, the sizes and diversity of sequence libraries are limited by the efficiency of transformation and by the nature of the protein to be studied. For example, if proteins are detrimental to cells, or have important regulatory functions within cells, they cannot be selected by these methods any more.
In the case of RIDS, a gene encoding a toxin that inactivates eukaryotic ribosomes, the ricin A chain (RTA) is fused downstream the sequence library to enable the complex formation. As a result, the functional proteins can be isolated under standard translation conditions without the need to remove the termination codon. RIDS is therefore perceived to be a powerful technique for the selection and evolution of peptides and proteins, useful for protein engineering researches including but not limited to:
• Evolution of Binding Agents
• Identification of Natural Protein–Protein Interactions
• Improvement of Protein Stability and Folding
• Specificity Profiling of Peptide-Binding Modules
• Mapping of Binding Energetics
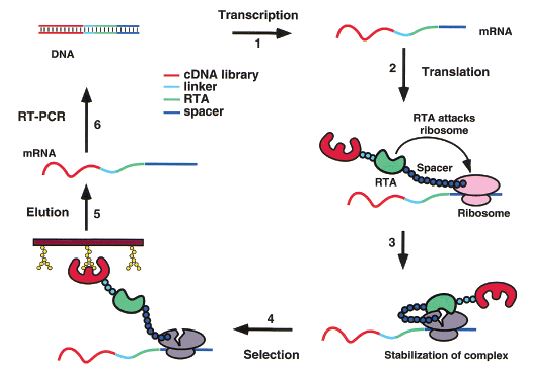
Figure 1: Schematic representation of the RIDS for screening functional proteins in vitro. (Jing-Min Zhou et al, 2002)
Reference
All listed services and products are For Research Use Only. Do Not use in any diagnostic or therapeutic applications.