Introduction of CHRNA10
CHRNA10, also known as neuronal acetylcholine receptor subunit alpha-10 (NACHRA10), is a protein that in humans is encoded by the CHRNA10 gene. Neuronal acetylcholine receptors (nAChRs), belonging to members of the Cys-loop family of ligand-gated ion, are pentameric receptors found both at the neuromuscular junction as well as in central and peripheral neurons. In vertebrates, nine nonmuscle alpha (α2–α10) and three betas (β2–β4) subunits have been identified. Different sets of α and β subunits result in the formation of nAChRs with different pharmacology and localization in the central and peripheral nervous system. The rules of association for functional nAChRs are broadening and now permit receptors assembled from single α subunits s (α7, α8, and α9) and receptors without supplemental subunits (α7α8, α9α10). The α10 subunit is a partner of α9 which is located in the inner hair cells and outer hair cells of the cochlea and the heteromeric α9α10 nAChR have important physiological functions in cochlear hair cells.
| Basic Information of CHRNA10 | |
| Protein Name | Neuronal acetylcholine receptor subunit alpha-10 |
| Gene Name | CHRNA10 |
| Aliases | Nicotinic acetylcholine receptor subunit alpha-10 |
| Organism | Homo sapiens (Human) |
| UniProt ID | Q9GZZ6 |
| Transmembrane Times | 4 |
| Length (aa) | 450 |
| Sequence | MGLRSHHLSLGLLLLFLLPAECLGAEGRLALKLFRDLFANYTSALRPVADTDQTLNVTLEVTLSQIIDMDERNQVLTLYLWIRQEWTDAYLRWDPNAYGGLDAIRIPSSLVWRPDIVLYNKADAQPPGSASTNVVLRHDGAVRWDAPAITRSSCRVDVAAFPFDAQHCGLTFGSWTHGGHQLDVRPRGAAASLADFVENVEWRVLGMPARRRVLTYGCCSEPYPDVTFTLLLRRRAAAYVCNLLLPCVLISLLAPLAFHLPADSGEKVSLGVTVLLALTVFQLLLAESMPPAESVPLIGKYYMATMTMVTFSTALTILIMNLHYCGPSVRPVPAWARALLLGHLARGLCVRERGEPCGQSRPPELSPSPQSPEGGAGPPAGPCHEPRCLCRQEALLHHVATIANTFRSHRAAQRCHEDWKRLARVMDRFFLAIFFSMALVMSLLVLVQAL |
Function of CHRNA10 Membrane Protein
The knowledge of the α10 nAChR distribution is still very limited, but surprisingly, it is coexpressed with the α9 subunit in rat and human mechanosensory hair cells. The α9 and α10 nicotinic cholinergic subunits assemble to form the receptor α9α10 nAChRs are believed to mediate synaptic transmission between efferent olivocochlear fibers and hair cells of the cochlea. The physiological role of α9α10 nAChRs in the auditory system has been determined by studies conducted with α9 and α10 knockout mice. The α9α10 nAChR was first identified in the auditory system where it mediates synaptic transmission between efferent olive cochlear cholinergic fibers and cochlear hair cells. This receptor has received further attention due to its potential role in chronic pain and breast and lung cancer. In human lymphocytes co-expressing the α9 and α10 subunit mRNAs, ACh is unable to produce ionic currents, indicating that the physiological effects of α9α10 nAChRs are different from those observed in cochlear hair cells.
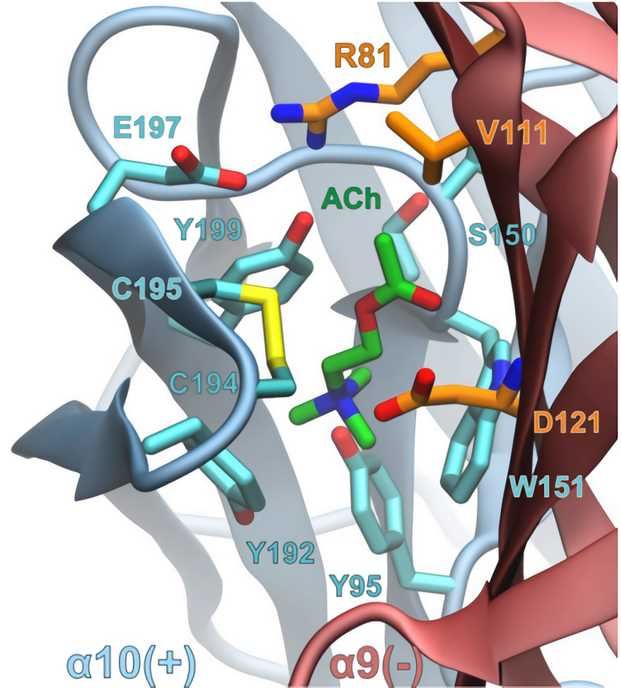 Fig.1. Close-up view of the α10/α9 binding interface from a representative structure of the molecular dynamics simulations of rat (α9)2(α10)3 complex with ACh. (Azam, 2015)
Fig.1. Close-up view of the α10/α9 binding interface from a representative structure of the molecular dynamics simulations of rat (α9)2(α10)3 complex with ACh. (Azam, 2015)
Application of CHRNA10 Membrane Protein Literature
This article uses α-CTx RgIA and acetylcholine to mimic rat α9α10 ECD and its complexes to demonstrate the interaction of α-CTx RgIA at α10/α9 but not α9/α10 nAChR subunit interface.
This article demonstrates that cyclization with a six-residue linker does not disturb the overall structure of RgIA, increases the selectivity for GABA(B) receptor over α9α10 nAChR, improve stability in human serum.
This article shows that dorsal root ganglion neurons coexpress α9 and α10 nicotinic receptor subunits.
This article describes the α9α10 nAChR may be a potential target in the pathophysiology of chronic pain, as well in the development of breast and lung cancers.
This article shows that the α9 and α10 nAChR subunits are expressed in different neurons and brain mitochondria of mouse brain and are compensatorily upregulated in the absence of the α7 subunit.
CHRNA10 Preparation Options
Membrane protein research has made significant progress over the past few years. Based on our versatile Magic™ membrane protein production platform, we could offer a range of membrane protein preparation services for worldwide customers in reconstitution forms and multiple active formats. Aided by our versatile Magic™ anti-membrane protein antibody discovery platform, we also provide customized anti-CHRNA10 antibody development services.
In the past years, Creative Biolabs has successfully produced many functional membrane proteins for our global customers. We are very honored to accelerate the development of our clients’ programs through our one-stop, custom-oriented service. For more detailed information, please feel free to contact us.
Reference
All listed services and products are For Research Use Only. Do Not use in any diagnostic or therapeutic applications.