CRISPR mediated Apoptosis related Gene Knockout Screening Service
Introduction
Creative Biolabs' CRISPR mediated Apoptosis-related Gene Knockout Screening Service uses advanced high-throughput lentiviral library design and functional screening in complex models. It systematically identifies novel pro-survival or pro-apoptosis genes, validates targets, and deciphers cancer drug resistance mechanisms. As a trusted functional genomics collaborator, we provide high-confidence candidates, expedite oncology drug discovery, and mitigate high attrition rates to enable next-generation precision therapeutics.
Discover How We Can Help - Request a Consultation Today
CRISPR mediated Apoptosis related Gene Knockout Screening Service
Background of Apoptosis-Related Genes
The balance of cell life and death is tightly controlled by families of pro-apoptotic and anti-apoptotic genes, most notably the BCL-2 family (e.g., BCL2, BIK, BBC3, BCLAF1, PERP) and the Caspases (e.g., CFLAR, CASP8). The dysregulation of these genes determines whether a cell succumbs to therapeutic stress or survives. Understanding the specific single-cell expression patterns of these regulators is essential to predict drug efficacy and resistance.
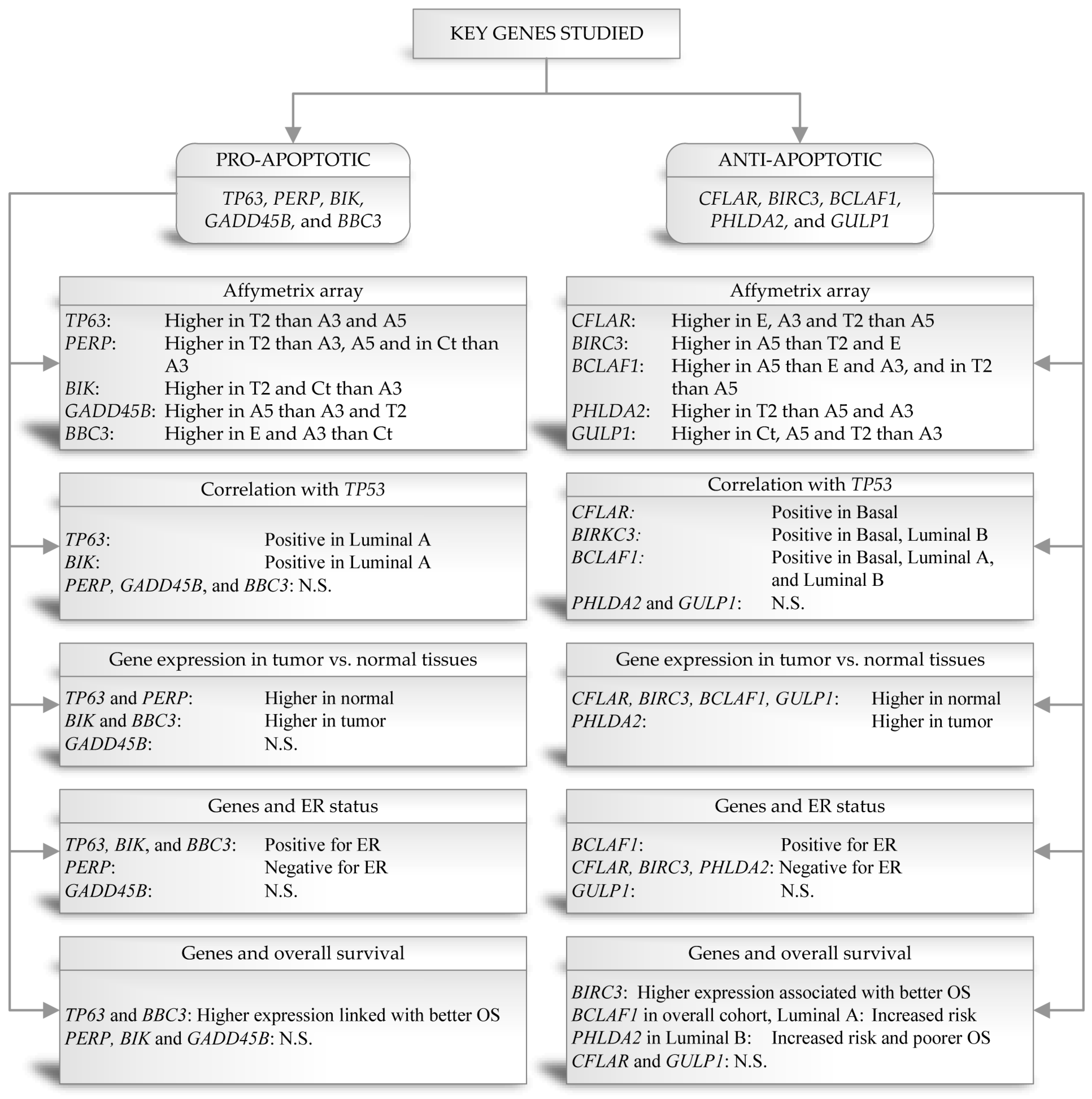 Fig.1 Genes related to apoptosis in experimental breast cancer models.1
Fig.1 Genes related to apoptosis in experimental breast cancer models.1
Screening Purpose
The primary purpose of the service is to systematically identify and validate genetic dependencies that modulate a cell's response to an apoptotic stimulus (e.g., a therapeutic compound or radiation). This functional screen moves beyond correlational data to pinpoint direct genetic requirements for survival or death in the context of your specific research questions.
Subsequent Application
Successful hits from the screen are immediately applicable to multiple downstream activities:
- Lead Optimization: Prioritizing drug candidates that show synthetic lethality with a known tumor vulnerability.
- Biomarker Development: Developing companion diagnostics based on the expression status of the identified hits to predict patient response.
- Combination Therapy Rationalization: Designing clinical trials that combine the lead compound with inhibitors targeting the newly identified resistance pathways.
Workflow
Our comprehensive workflow is designed for speed, clarity, and biological relevance, ensuring a smooth transition from project initiation to final results.
| Aspect | Description of Activities & Expected Outcomes |
|---|---|
| Required Starting Materials |
1. Target Gene List/Pathway Focus: A list of 50-500 candidate apoptosis genes or a specific disease pathway to screen. 2. Cell Line/Model: Your chosen cancer cell line, patient-derived organoids (PDOs), or specialized co-culture model. 3. Treatment Conditions: Concentration and time-point range for the therapeutic agent(s) of interest. |
| Lentiviral Library Engineering & gRNA Fabrication | We develop a high-accuracy, refined gRNA library (pooled or arrayed) directed at your target genes, guaranteeing optimal specificity and coverage. |
| Virus Production & Cell Transduction | High-titer lentivirus is produced and optimized for maximum cell transduction efficiency (low MOI) in your specific model to ensure single-copy integration. |
| Targeted Intervention & Deep Sequencing | Transduced cells are exposed to the designated treatment (e.g., drug, radiation). |
| Bioinformatics Analysis & Hit Identification | Leveraging sophisticated statistical models, we conduct rigorous normalization and differential enrichment analysis to identify high-confidence "hit" genes that exert a significant impact on cell survival. |
| Lead Candidate Validation | Top hit genes are individually validated using secondary assays (e.g., viability, colony formation) to confirm the screening results and eliminate false positives. |
| Final Deliverables |
1. Gene Hit List Report: A prioritized table of genes ranked by significance, p-value, and fold-change for sensitization and resistance. 2. Raw and Processed Sequencing Data: Fully annotated FastQ files and count matrices. 3. Cell-Specific Survival Curve Report: Graphical and quantitative validation of top candidates in your cell model. |
| Estimated Timeframe | The typical timeframe for this service ranges from 8 to 12 weeks, depending on the complexity of the cell model (e.g., PDOs often require longer optimization) and the size of the target library. |
What We Can Offer
Creative Biolabs is your expert partner, providing fully customizable CRISPR screening solutions to meet the most stringent demands of modern oncology research. Our technical offerings ensure that your functional genomics project delivers high-quality, actionable data every time.
Tailored gRNA Library Design
Expert, custom design of focused apoptosis gene libraries (50–500 targets) or whole-genome scale libraries, ensuring optimal coverage and minimizing off-target effects specific to your research objectives.
Unmatched Model Flexibility
Proven technical proficiency in running screens across challenging, translationally relevant systems, including patient-derived organoids (PDOs), 3D co-culture models, and primary immune cells, accurately mimicking the in vivo environment.
High-Titer Lentiviral Delivery
Optimized, large-scale lentivirus production ensures maximum transduction efficiency (high quality, low multiplicity of infection/MOI) and reliable, single-copy gRNA integration across diverse cell types.
Robust Bioinformatics & Data Quality
Utilization of cutting-edge statistical models for hit-calling, normalization, and batch-effect correction, adhering to principles of Quality-by-Design (QbD) to guarantee the integrity and confidence of your final Gene Hit List.
Comprehensive Validation Support
Integrated follow-up services for individual gene validation using orthogonal assays (e.g., qPCR, Western Blot, customized viability assays) to transition candidates quickly from screen to functional proof-of-concept.
Experience the Creative Biolabs Advantage - Get a Quote Today
Customer Reviews

FAQs
How does Creative Biolabs' CRISPR screening compare to traditional shRNA screens for apoptosis?
CRISPR-Cas9 provides a virtually complete gene knockout, leading to stronger phenotypic effects and fewer off-target effects compared to shRNA, which only achieves knockdown. Our optimized gRNA libraries and rigorous data analysis minimize false positives, giving you higher confidence in your hit list.
We use patient-derived organoids (PDOs). Can your service handle these complex, low-passage models?
Absolutely. Our expertise lies in optimizing transduction protocols (low MOI) and ensuring cell viability in delicate models like PDOs and primary cells. This capability is key to generating data that is highly translatable to the clinic.
Our therapeutic agent causes high initial toxicity. How can we ensure we get meaningful hits?
We work with you to design a tailored screen format, including multi-dose selection and rescue time-points, which helps distinguish true resistance mechanisms from general cell death. This maximizes the biological signal and ensures we identify only the most relevant genetic modulators.
What if we don't have a final list of target genes but only a general idea (e.g., "Mitochondrial Apoptosis")?
That's fine. Our expert scientific team can assist you in curating a highly relevant, focused library of apoptosis-related genes based on literature and public domain data (TCGA, GTEx) specific to your disease area, ensuring the screening effort is highly strategic.
Creative Biolabs offers the industry-leading CRISPR mediated Apoptosis related Gene Knockout Screening Service, providing the high-resolution functional validation required to accelerate your oncology pipeline. From initial library design through to actionable final Gene Hit Lists, we stand as your committed collaborator in translating genomic complexity into therapeutic achievement.
Contact Our Team for More Information and to Discuss Your Project
Reference
- Calaf, Gloria M., and Leodan A. Crispin. "Genes Associated with Apoptosis in an Experimental Breast Cancer Model." International Journal of Molecular Sciences 26.19 (2025): 9735. https://doi.org/10.3390/ijms26199735. Distributed under Open Access license CC BY 4.0, without modification.
