CRISPR mediated Kinase Knockout Screening Service
Introduction
Creative Biolabs' CRISPR mediated Kinase Knockout Screening Service uses CRISPR-Cas9 to systematically disrupt human kinase genes (≈500), identifying those essential for cellular functions. It addresses post-transcriptional controls (e.g., Alternative Polyadenylation) and uses rigorous computation for reliable, clinical data. The service accelerates drug discovery via high-throughput precision engineering, delivers actionable data on essential kinase targets, and moves projects from hypothesis to validated targets for downstream development.
Discover How We Can Help - Request a Consultation Today
CRISPR Mediated Kinase Knockout Screening Service
Background of Kinase Screening
Kinases are responsible for transferring phosphate groups from ATP to target proteins, thereby regulating nearly every cellular process. Due to their central regulatory role, approximately 25% of current drug discovery programs target kinases. Traditional screening relied on siRNA or chemical libraries, which often suffer from off-target effects, inconsistent knockdown, and high cost. CRISPR-Cas9 offers a superior, permanent, and highly specific gene knockout that perfectly models a functional loss-of-function phenotype for target validation.
Screening Purpose
The primary purpose of the CRISPR mediated Kinase Knockout Screen is to move from correlation to causation. Specifically, it aims to:
- Identify Novel Targets: Systematically reveal previously unrecognized kinases essential for a disease state or cellular function (e.g., identifying a metabolic regulator like EXOSC10 as a key vulnerability).
- Validate Existing Targets: Confirm the functional relevance of known or suspected kinase hits with high confidence.
- Determine Resistance Mechanisms: Identify kinases whose loss confers resistance or sensitivity to a specific compound, informing combination therapy strategies.
Subsequent Application
The high-confidence data generated by this service directly drives several downstream applications:
- Drug Discovery: The validated kinase target is immediately prioritized for small-molecule inhibitor design, antibody development, or PROTAC degradation.
- Biomarker Identification: The screen identifies genetic signatures that can be used for patient stratification in clinical trials (similar to using a genetic profile for prognosis).
- Process Optimization: For biomanufacturing clients, the identified targets can be used to engineer production cell lines with enhanced growth, improved viability, or higher product titer.
Workflow
Our workflow is designed for maximum efficiency and clarity, suitable for visualization as a flowchart, ensuring clients understand every stage of the process. The typical timeframe for this service ranges from 8 to 14 weeks, depending on the complexity of the cell model and the depth of the subsequent validation required.
| Stage | Description of Activities Involved |
|---|---|
| Project Initiation & Design | Required Starting Materials: Target Cell Line(s) (e.g., specific cancer lines, iPSC derivatives), Defined Selection Criteria (e.g., viability under drug X, proliferation rate, expression of biomarker Y), and Preliminary Data (e.g., initial hit lists, pathway hypotheses). |
| Transduction & Selection | Lentiviral packaging and high-efficiency transduction of the custom sgRNA library into the client's cell line. Cells are then subjected to the defined selective pressure (e.g., drug treatment, specific media conditions) over a precise time course. |
| High-Throughput Sequencing & Data Analysis | Genomic DNA extraction from selected cell pools. High-throughput next-generation sequencing (NGS) of the integrated sgRNA sequences to determine their relative abundance. Computational analysis using robust algorithms (e.g., MAGeCK) to calculate the Guide RNA Log-Fold Change (LFC) scores. |
| Validation & Secondary Screening | Single-gene knockout cell line generation for the top 5-10 hits from the primary screen. These individual lines are re-tested using the original selective criteria (e.g., IC50 shift) to confirm the primary findings and eliminate false positives. |
Final Deliverables:
- Comprehensive Screening Report: Detailed NGS data, raw read counts, LFC scores, and statistical analysis for every kinase in the library.
- Validated Target List: A summary of the top-performing kinase genes with supporting functional data from secondary validation assays.
- Engineered Cell Line(s): Optional delivery of the final single-knockout cell lines generated during the validation stage.
What We Can Offer
Precision and Specificity
We utilize advanced sgRNA design algorithms and robust computational analysis to specifically target the human kinome, providing unprecedented functional clarity while minimizing off-target effects. This is a critical advantage over traditional knockdown methods (siRNA).
Customized Library Design
Beyond the standard human kinome, we offer fully customizable sgRNA library synthesis. It integrates specific pathways, synergistic targets (dual gRNA screens), or specific gene families relevant to your unique research focus.
Integrated DBTL Cycle Support
We go beyond the "Test" phase by offering seamless integration into the Design, Build, and Learn cycle. This includes the subsequent generation of validated single-knockout cell lines for your top hits, accelerating your preclinical development.
Advanced Computational Validation
Our service includes high-fidelity NGS data analysis (LFC scoring) backed by sophisticated statistical models similar to those used in high-impact prognostic studies. This ensures your hit list is based on statistically sound, high-confidence data.
High-Standard Quality Control
Strict adherence to robust QC standards throughout library production, transduction, and NGS sequencing ensures the output is not just data, but a validated, high-quality target set ready for lead identification and optimization.
Case Study
To systematically identify the regulatory factors of protein kinases in the TGF-β signaling pathway, CRISPR-Cas9 screening was conducted in HeLa cells using the human whole-kinase group CRISPR-Cas9 knockout library. After transfection for 48 hours, puromycin was added to screen the transfected cells and continued to culture for 48 hours. 2× 107 cells were collected as pre-screening cells to evaluate the library integrity and homogeneity of samples after lentivirus transduction. The remaining cells were collected after culturing for 7 days as pre-sorting cells. HeLa cells were stimulated with TGF-β1 for 1 hour, and the cells with active and inactive TGF-β signaling pathways were sorted according to the expression level of P-Smad2. Further sequencing work will be carried out.
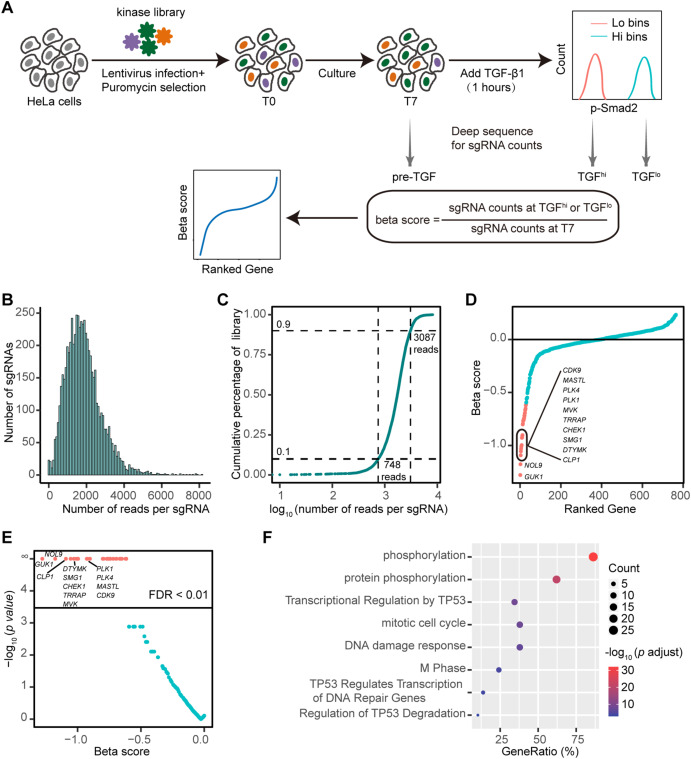 Fig.1 Full kinase group CRISPR-Cas9 knockout screening was performed in HeLa cells to identify the regulatory factors of TGF-β signaling.1
Fig.1 Full kinase group CRISPR-Cas9 knockout screening was performed in HeLa cells to identify the regulatory factors of TGF-β signaling.1
Customer Reviews

Experience the Creative Biolabs Advantage - Get a Quote Today
FAQs
How does CRISPR screening compare to traditional siRNA screening for target identification?
CRISPR-Cas9 provides a clean, permanent genetic knockout, whereas siRNA only offers transient knockdown, which can be inconsistent. Our CRISPR method offers higher confidence in the functional relevance of the target, leading to more robust and predictive data for downstream drug development.
What input material is necessary, and do you handle complex or primary cell lines?
We are experts in handling challenging cell models. We require your target cell line and a clear, measurable selection criterion. Yes, our high-efficiency lentiviral transduction protocols are optimized for complex models, including iPSCs and some primary cells.
Can your service identify synergistic or synthetic lethal relationships between kinases?
Absolutely. While the standard kinome screen targets single genes, we also offer custom pooled dual-sgRNA libraries designed to probe interactions between two different kinase genes or between a kinase and a CRISPR editor, which is essential for defining combination therapy strategies.
We are concerned about off-target effects. How does Creative Biolabs ensure data specificity?
We address this with multiple layers of quality control. We utilize advanced sgRNA design algorithms to select guides with minimal predicted off-target activity, and our computational analysis (using LFC scores) employs statistical methods to confidently differentiate true hits from background noise, which we then validate with single-gene knockout assays.
Creative Biolabs' CRISPR mediated Kinase Knockout Screening Service delivers a definitive, high-confidence solution for functional target validation. By integrating cutting-edge genomic tools with expert computational analysis, we transform long, unpredictable discovery cycles into a streamlined, results-driven process, ensuring your project moves rapidly from hypothesis to clinically relevant action.
Contact Our Team for More Information and to Discuss Your Project Today
Reference
- Wang, Dingding, et al. "Kinome-wide CRISPR-Cas9 screens revealed EXOSC10 as a positive regulator of TGF-β signaling." Biochemistry and Biophysics Reports 40 (2024): 101864. https://doi.org/10.1016/j.bbrep.2024.101864. Distributed under Open Access license CC BY 4.0, without modification.
