CRISPR mediated Protein Homeostasis Knockout Screening Service
Introduction
Creative Biolabs' CRISPR mediated Protein Homeostasis Knockout Screening Service uses genome-wide CRISPR-Cas9 and high-fidelity phenotypic readouts. It maps PQC failure's genetic architecture, identifies novel regulatory factors, and validates causal targets linked to toxic protein aggregation. As a trusted partner, we deliver mechanistic insights, de-risk drug pipelines by focusing on key PQC pillars, and accelerate discovery for neurodegenerative disease therapeutics.
CRISPR mediated Protein Homeostasis Knockout Screening Service
Background of Protein Homeostasis
Protein Homeostasis (PQC) represents the cell's integrated defense system against proteotoxicity. It is essential for managing misfolded and damaged proteins that accumulate in neurodegenerative diseases like ALS and Alzheimer's. This system comprises three interacting branches: Molecular Chaperones (to assist folding), the Ubiquitin-Proteasome System (UPS, for selective, immediate degradation), and the Autophagy-Lysosomal System (for bulk degradation of large aggregates like aggresomes). Age-related decline compromises all three pillars, necessitating therapeutic strategies that specifically enhance the function of these failing systems.
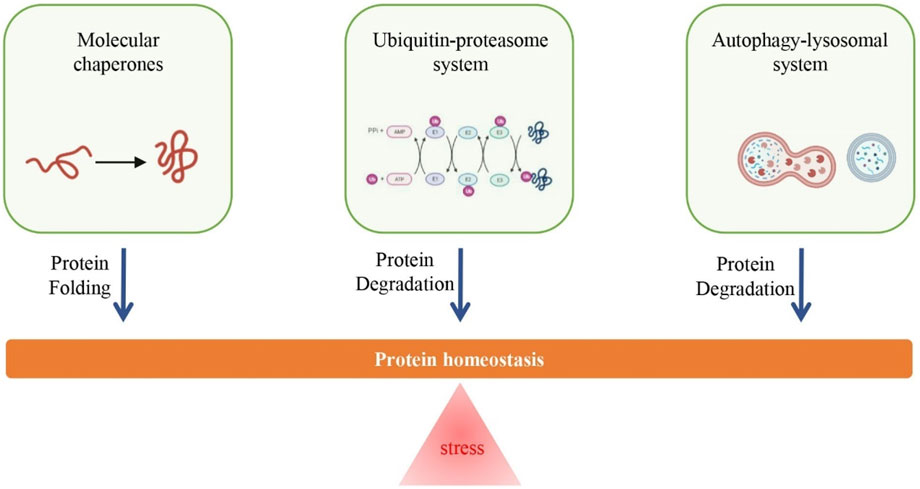 Fig.1 Long-term chronic stress is an important cause of protein homeostasis loss, but the enhancement of the ubiquitin-proteasome system, autophagy-lysosomal system, and molecular chaperone function can promote the maintenance of protein homeostasis.1,3
Fig.1 Long-term chronic stress is an important cause of protein homeostasis loss, but the enhancement of the ubiquitin-proteasome system, autophagy-lysosomal system, and molecular chaperone function can promote the maintenance of protein homeostasis.1,3
Screening Purpose
The primary purpose of the Creative Biolabs CRISPR mediated Protein Homeostasis Knockout Screening Service is to move beyond mere correlation and identify causal genetic modifiers that directly enhance or inhibit the PQC response to aggregation-prone proteins. By systematically knocking out every non-essential gene, we uncover both known and novel components of the PQC network, providing a comprehensive map of the genes that govern protein folding, stability, and clearance in a disease-relevant cellular context.
Subsequent Application
The high-confidence targets generated by our screening service are immediately applicable to multiple stages of the drug discovery pipeline. The validated targets can be moved directly into High-Throughput Screening (HTS) campaigns to search for small molecule modulators that mimic the protective genetic knockout phenotype. They also serve as the foundation for Mechanistic Analysis, including proteomics and advanced imaging, which defines the drug mechanism of action. Furthermore, the genetic insights can inform the development of novel gene therapies, such as AAV-mediated overexpression of protective DUBs or PQC-enhancing factors.
Workflow
Required Starting Materials
To initiate the service, a client typically provides:
- Disease-Relevant Cellular Model: e.g., iPSC-derived neurons expressing a known aggregation-prone mutant protein (e.g., TDP-43 M337V, Tau P301L).
- Reporter System Documentation: Details on the assay or readout already established to detect protein aggregation (e.g., fluorescently tagged inclusion bodies).
- Project Scope & Target Criteria: Clear specifications on the desired gene set (whole genome or focused library) and hit-selection criteria.


Model Optimization and Library Generation
Creative Biolabs engineers the client's cellular model to ensure robust and stable expression of the aggregation phenotype. We then transduce cells with a high-complexity, genome-wide CRISPR-Cas9 knockout library to ensure every human gene is systematically interrogated.
High-Content Phenotypic Screening
Cells are subjected to the genome-wide knockout and grown under defined stress conditions. We use high-content imaging and FACS-based sorting methods, such as the Pulse Shape Analysis (PulSA) approach, to achieve direct, quantitative detection of protein inclusion formation at the single-cell level.


Genetic Hit Identification and Deconvolution
Following the screen, the specific gRNA sequences enriched (genes that reduce aggregation) or depleted (genes that increase aggregation) are analyzed using next-generation sequencing. This step unmasks the top causal modifier genes.
Target Validation and Functional Triage
Individual candidate genes are subjected to secondary screening using single gRNAs to confirm the knockout phenotype. Functional assays (e.g., western blot, solubility assays, microscopy) are performed to establish the mechanistic role of the novel target in PQC.


Final Report and Data Delivery
A comprehensive report detailing the top-tier hits, their validation data, proposed mechanism of action, and a prioritized list of targets suitable for follow-on studies is provided.
Final Deliverables
Upon completion, the client receives:
- Prioritized Target List: A ranked list of 10-20 high-confidence genes validated as causal modifiers of proteostasis failure.
- Mechanistic Data Package: Detailed functional data, including microscopy images of aggregate modulation, and quantitative data on protein solubility and stability.
- Sequencing Data and Analysis Report: Raw and processed sequencing data from the CRISPR screen for internal analysis and future reference.


Estimated Timeframe
The typical timeframe for this service ranges from 16 to 24 weeks, depending on the complexity of the cellular model provided and the scope of the screen (whole-genome vs. focused library).
Discover How We Can Help - Request a Consultation
What We Can Offer
Creative Biolabs' CRISPR mediated Protein Homeostasis Knockout Screening Service offers biology experts a highly resolved, genetically driven platform to solve the most difficult challenges in neurodegenerative disease. We emphasize technical excellence and complete project customization.
Our Advantage
Mechanistic Clarity
Genome-wide, unbiased screening systematically interrogates the function of all human genes to provide causal targets, eliminating false positives from general cell toxicity.
Precision Phenotypic Readouts
Deployment of proprietary high-fidelity methods like the Pulse Shape Analysis (PulSA) for direct, quantitative detection of protein inclusion formation at the single-cell level.
Comprehensive PQC Coverage
Targeted screening across the entire Protein Quality Control (PQC) network, including Molecular Chaperones, the Ubiquitin-Proteasome System (UPS) (e.g., DUB/E3 identification like BAP1), and the Autophagy-Lysosomal System (e.g., structural components like SRRD).
Unmatched Customization
Complete flexibility in adapting the platform to your specific aggregation-prone protein (TDP-43, Tau, α-synuclein, Huntingtin) and disease-relevant cellular model (iPSC-derived neurons, glia, etc.).
Actionable Deliverables
Data-driven, prioritized targets that have been functionally validated to directly modulate the desired PQC failure mechanism, ready for immediate HTS or lead optimization.
Scientific Partnership
Deep expertise from the Creative Biolabs team to design the optimal gRNA library, select stress conditions, and interpret complex sequencing data to maximize discovery impact.
Case Study
Genome-wide CRISPR-Cas9 knockout screening was conducted to identify the gene modification factors for the formation of TPP-43 ΔNLS protein inclusion bodies in human cells. In addition to the known genes related to cellular protein homeostasis, our CRISPR KO screening also identified some interesting inclusion body formation modification genes. One of the genes is METTL5, a newly discovered 18S rRNA m6A methyltransferase that plays a role in translation initiation and is a regulator of cellular stress response. Another interesting gene is XPO4, a nuclear export factor. When it is absent, the number of cells containing McLover3-tpp-43 Δ NLS aggregates will also increase.
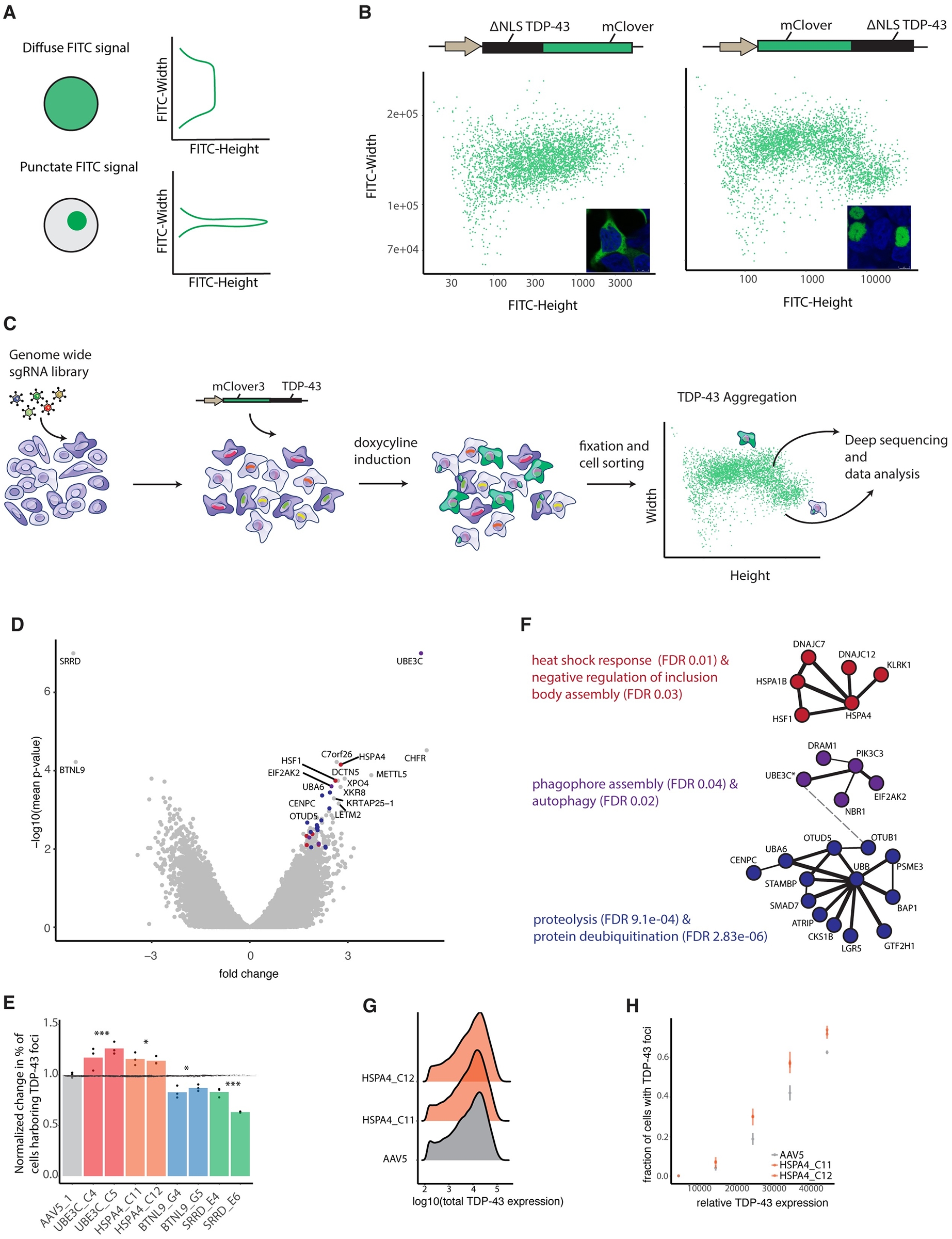 Fig.2 Genome-wide CRISPR-Cas9 knockout screening leveraging impulse shape analysis uncovered both anticipated and previously unreported regulators of cellular protein homeostasis.2,3
Fig.2 Genome-wide CRISPR-Cas9 knockout screening leveraging impulse shape analysis uncovered both anticipated and previously unreported regulators of cellular protein homeostasis.2,3
Customer Reviews

Experience the Creative Biolabs Advantage - Get a Quote Today
FAQs
How does this CRISPR screen compare to a standard chemical screen for aggregation inhibitors?
Standard chemical screens rely on libraries of compounds that may hit non-specific or toxic targets, often revealing general toxicity effects rather than causal mechanisms. Our CRISPR screen provides a genetic map of causality, revealing which gene is responsible for the effect. Once a causal gene is identified (e.g., SRRD or BAP1), the subsequent chemical screen is far more focused and mechanistic, leading to higher quality, less toxic leads.
What if our target protein aggregates are too rare or unstable to screen effectively?
We utilize highly sensitive, single-cell analysis methods like FACS and high-content imaging, often leveraging fluorescent reporters developed for high-fidelity detection. This allows us to quantify subtle changes in aggregation dynamics that may be missed by bulk assays. We work closely during the Model Optimization stage to ensure the assay is robust before launching the genome-wide screen.
Can your platform target components of the Ubiquitin-Proteasome System (UPS) like DUBs?
Absolutely. As shown by the BAP1/SPIN4 example, DUBs are high-value targets. By focusing on the direct regulation of target protein stability, our knockout screens can precisely identify DUBs and E3 ligases that control the degradation fate of aggregation-prone proteins, providing powerful, enzymatic targets for stabilization or clearance.
Our therapeutic strategy requires a non-toxic approach. Does this platform inherently favor non-toxic hits?
Yes. By specifically measuring the formation or clearance of protein aggregates as the primary readout (phenotype) rather than general cell death (toxicity), our platform inherently selects for genes that modulate the underlying mechanism. These mechanistic targets enable therapeutic strategies that restore function, which are typically less toxic than those that merely suppress symptoms.
Creative Biolabs' CRISPR mediated Protein Homeostasis Knockout Screening Service is the definitive platform for de-risking and accelerating drug discovery in neurodegenerative disease. We transform the complex failure of proteostasis into a clear set of actionable genetic targets, using precision tools to ensure your next therapeutic candidate is built upon a foundation of causality.
Contact Our Team for More Information and to Discuss Your Project
References
- Chen, Xiao-Qiong, et al. "Protein homeostasis in aging and cancer." Frontiers in cell and developmental biology 11 (2023): 1143532. https://doi.org/10.3389/fcell.2023.1143532.
- Sweeney, Katelyn M., et al. "CRISPR screen for protein inclusion formation uncovers a role for SRRD in the regulation of intermediate filament dynamics and aggresome assembly." PLoS Genetics 20.2 (2024): e1011138. https://doi.org/10.1371/journal.pgen.1011138.
- Distributed under Open Access license CC BY 4.0, without modification.
