CRISPR mediated Cell Cycle Protein Knockout Screening Service
Introduction
Genomic instability from DNA Damage Response (DDR) pathway defects is a key cancer characteristic. Creative Biolabs' CRISPR mediated Cell Cycle Protein Knockout Screening Service uses advanced functional genomics and high-throughput CRISPR/Cas9 to systematically map gene dependencies, converting genetic associations into actionable therapeutic targets. It accelerates cancer drug discovery, identifies synthetic lethal vulnerabilities, enables high-confidence validation of DDR and cell cycle-related targets, and dissects cancer cell survival regulatory mechanisms.
Discover How We Can Help - Request a Consultation
CRISPR Mediated Cell Cycle Protein Knockout Screening Service
Background of Cell Cycle Proteins
Cell cycle proteins are the central machinery regulating cellular proliferation, DNA replication fidelity, and genome maintenance. This network includes key components responsible for checkpoint signaling (G1, S, G2, M phases) and DNA repair pathways (e.g., Homologous Recombination, Non-Homologous End Joining). In oncology, these proteins are frequently mutated or deregulated, making cancer cells heavily reliant on remaining checkpoint function for survival. Inhibiting these residual functions is the core principle behind synthetic lethality.
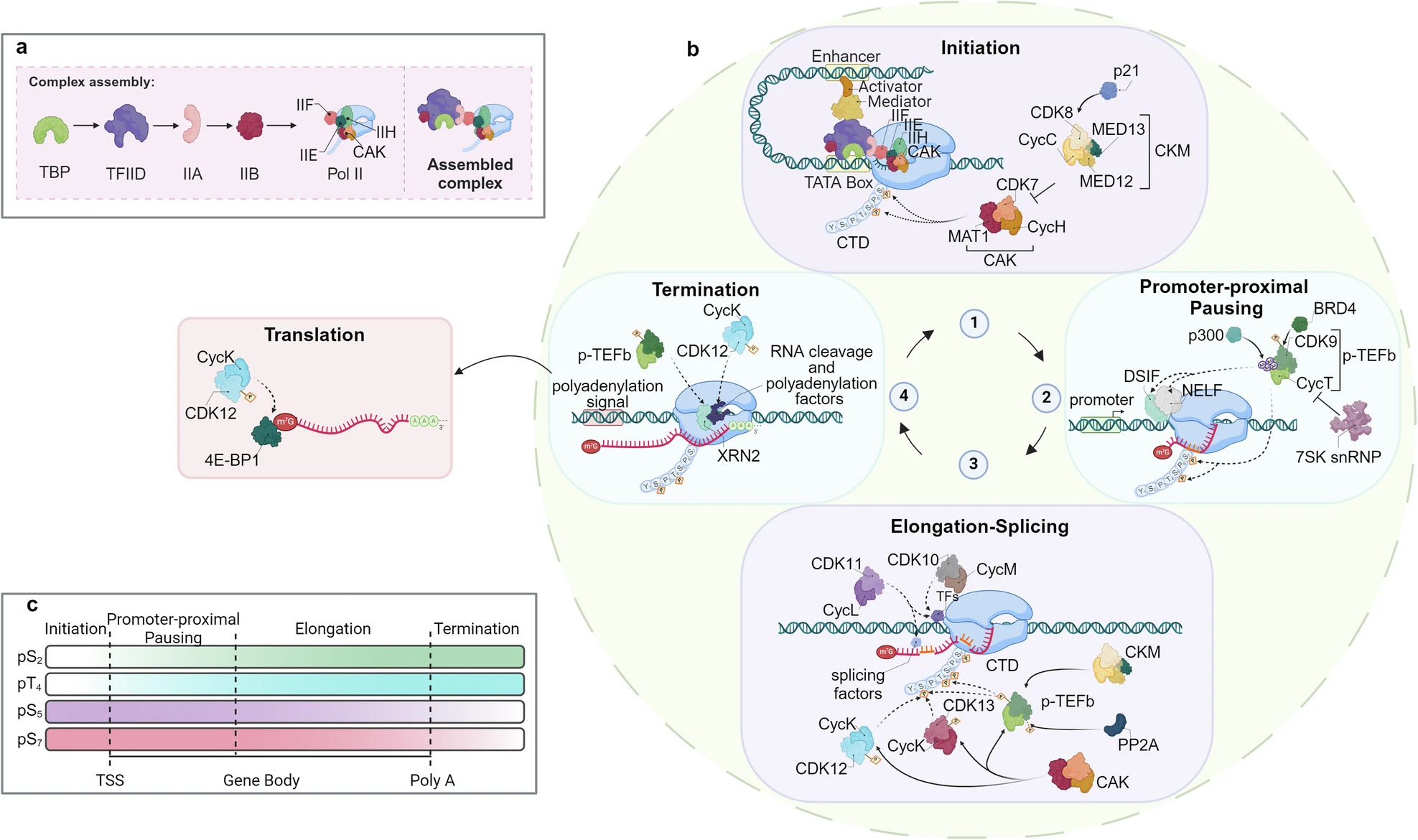 Fig.1 The role of CDK/ cyclin complexes in the regulation of the transcriptional cycle.1,3
Fig.1 The role of CDK/ cyclin complexes in the regulation of the transcriptional cycle.1,3
| Cyclin Family | Core Members | Binding CDK Partners | Primary Stage of Action | Core Functions |
|---|---|---|---|---|
| Cyclin D | D1, D2, D3 | CDK4, CDK6 | G1 phase | Respond to growth factor signals, phosphorylate Rb protein to release E2F transcription factors, drive cells from G0 phase to G1 phase, and initiate G1 phase progression. |
| Cyclin E | E1, E2 | CDK2 | Late G1 phase → G1/S transition | Further phosphorylate the Rb protein, activate the expression of DNA replication-related genes, promote cells to pass the G1/S checkpoint, and initiate S phase. |
| Cyclin A | A1, A2 | CDK2 (S phase), CDK1 (G2/M phase) | S phase → G2/M phase | Cyclin A2-CDK2: Maintain S phase DNA synthesis; Cyclin A2-CDK1: Regulate G2 phase progression and prepare for M phase. |
| Cyclin B | B1, B2 | CDK1 (Cdc2) | G2 phase → M phase transition | Form maturation-promoting factor (MPF), phosphorylates nuclear envelope proteins and chromosome condensation-related proteins, triggers nuclear envelope breakdown and chromosome condensation, and initiates mitosis. |
| Cyclin C | C | CDK8, CDK19 | G1 phase, transcriptional regulation | Combine with CDK8/19 to participate in transcriptional co-regulation; assist in early G1 phase regulation and the balance between cell proliferation and differentiation when bound to CDK3 or acting alone. |
| Cyclin F | F | None (does not bind CDK) | G2/M phase | It acts as an F-box protein to participate in the ubiquitin-proteasome pathway, degrade regulatory factors such as Cyclin A and Wee1, and maintain the homeostasis of the G2/M checkpoint. |
Tab.1 Cyclin-CDK Complex Function Comparison Table.
Subsequent Application
The high-confidence targets identified through this service have immediate downstream applications in translational oncology and drug development:
- Drug Target Prioritization: Validating the most promising molecular targets for small molecule inhibitor development (e.g., focusing on proteins that drive G2-M arrest, such as PSMD4 in CRC models).
- Predictive Biomarker Development: Utilizing identified hits (like components of the DDR pathway) to create more accurate biomarkers for patient selection and monitoring treatment efficacy.
- Novel Combination Therapies: Designing rational combination regimens where an identified target inhibitor is paired with an existing agent, creating enhanced therapeutic efficacy and overcoming drug resistance.
Workflow
Our comprehensive workflow is designed for maximum clarity and scientific rigor, making the transition from concept to validated target seamless for biopharma clients.
Required Starting Materials
- Specific Cancer Cell Line/Model: Your preferred cell line (e.g., breast, colon, lung) or patient-derived xenograft (PDX) model for translational relevance.
- Custom gRNA Library Design Specifications: Information on your gene set of interest (e.g., 200 DDR genes) or genome-wide library preference.
- Compound-of-Interest (CoI) Data: Preliminary dose-response and toxicity data for the drug used to select the surviving population.


gRNA Library Transduction and Selection
High-titer viral preparation of the custom gRNA library is used to transduce target cells. Outcome: A genetically diverse pool of cells, each lacking a specific target gene.
Application of Selective Pressure
Cells are treated with the CoI (e.g., a DDR inhibitor) at a defined concentration, enriching for cells that are either resistant or hypersensitive. Outcome: Elimination of non-essential cells and enrichment of desired phenotypes.


Genomic DNA Extraction and PCR
Genomic DNA is extracted from the surviving cell populations and the initial control pool. The integrated gRNA sequences are amplified. Outcome: Purified DNA template ready for quantification.
Next-Generation Sequencing (NGS) & Data Analysis
Amplified gRNA sequences are quantified via NGS, and enrichment/depletion ratios are calculated using sophisticated bioinformatics pipelines (e.g., MAGeCK). Outcome: A raw count table and statistically significant hit list (MAGeCK score >2.0).


Target Validation
Single-gene knockout cell lines for top candidates (e.g., PSMD4) are generated and validated using orthogonal assays (e.g., colony formation, cell cycle analysis). Outcome: Confirmation of the hit's functional role in sensitization or resistance.
Final Deliverables
- Validated Hit List (Ranked): A statistically significant, ranked list of genes influencing cellular fitness under treatment.
- Raw NGS Count Files: Complete raw and normalized sequencing data for internal client validation.
- Comprehensive Mechanism-of-Action (MOA) Report: Detailed analysis of the potential pathways affected, including cell cycle, apoptosis, and DDR involvement.


Estimated Timeframe
The typical timeframe for this service ranges from 12 to 16 weeks, depending on the complexity of the screening model, the size of the gRNA library (custom versus genome-wide), and the required depth of validation.
What We Can Offer
Creative Biolabs is the partner of choice for accelerating the identification of high-value targets in oncology. Our CRISPR mediated Cell Cycle Protein Knockout Screening Service is delivered with comprehensive capabilities and customization at every stage:
Tailored Screening Campaigns
Fully customizable gRNA library design, ranging from focused, pathway-specific panels (e.g., 200 DDR genes) to complete genome-wide arrays, perfectly optimized for your specific research question and cancer model.
Precision Functional Mapping
Dedicated high-resolution CRISPR/Cas9 functional screening specifically designed to map synthetic lethal vulnerabilities within the DDR and cell cycle checkpoint pathways.
High-Confidence Target Validation
Integration of NGS readout with industry-standard, robust bioinformatics pipelines (e.g., MAGeCK analysis) to deliver statistically validated hits and ranked target lists for immediate prioritization.
Translational Focus on Complex Biology
Unique expertise at Creative Biolabs in modeling complex, clinically relevant genotoxic events (including bacterial or environmental stressors) to uncover novel and impactful therapeutic dependencies.
End-to-End Quality Assurance
Our well-established quality system ensures maximum data reproducibility, providing detailed Mechanism-of-Action (MOA) reporting and streamlined project management from initial design to final validated targets.
Orthogonal Validation and Deliverables
Complete service package including downstream single-target validation using orthogonal assays (e.g., flow cytometry for cell cycle analysis, colony formation) to confirm the functional role of the hit in your model.
Experience the Creative Biolabs Advantage - Get a Quote Today
Case Study
Researchers used the GeCKO v2 sgRNA library to conduct genome-wide CRISPR/Cas9 gene knockout screening in HEK293T cells, aiming to identify the key genes that confer E. coli sensitivity on the cells. The results showed that the proteasome 26S subunit ubiquitin receptor (PSMD4) was significantly enriched under various screening conditions. Further verification revealed that the survival rates of both PSMD4 knockout HEK293T cell populations after treatment with Escherichia coli were significantly higher than those of wild-type cells. It is known that G2-M phase cell cycle arrest is a typical stress response of cells to Escherichia coli poisoning. Based on the above screening and verification results, researchers proposed a hypothesis: PSMD4 knockout may regulate cell survival by affecting the G2-M phase arrest process. Subsequent experiments confirmed that PSMD4 knockout significantly altered the proportion of G2-M phase blocked cells, providing crucial support for the mechanism of E. coli resistance mediated by it.
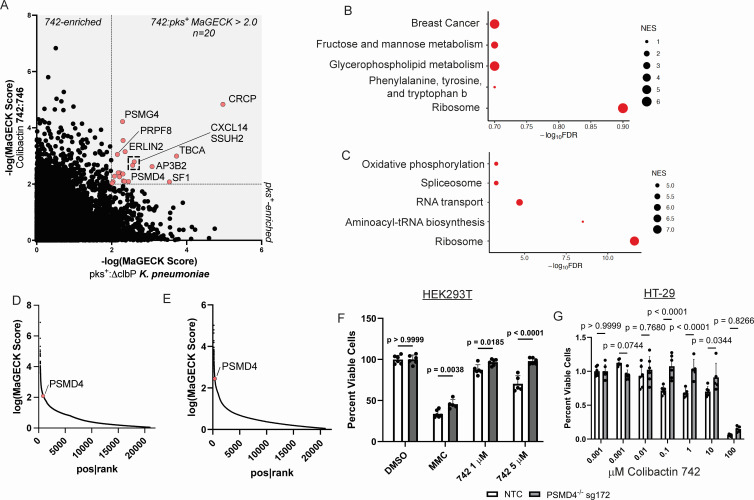 Fig.2 After screening the whole genome of Escherichia coli using CRISPR KO, sequencing was performed, and the significantly enriched genes were selected for further analysis.2,3
Fig.2 After screening the whole genome of Escherichia coli using CRISPR KO, sequencing was performed, and the significantly enriched genes were selected for further analysis.2,3
Customer Reviews

FAQs
Can Creative Biolabs customize the screening to focus on specific DDR pathways, or must it be genome-wide?
We specialize in custom solutions. While a genome-wide screen offers the broadest coverage, we frequently design focused libraries targeting specific DDR sub-pathways (e.g., Homologous Recombination, Fanconi Anemia) or cell cycle regulators. This strategy significantly reduces screening time and costs while maintaining high functional resolution.
How do you ensure the high quality and specificity of the knockout data?
Our stringent quality control measures include multi-point gRNA library deep sequencing, extensive biological replication, and advanced bioinformatics algorithms (like MAGeCK) to distinguish true hits from noise. We also recommend single-target validation assays (e.g., flow cytometry, colony formation) to confirm that the observed phenotype is genuinely linked to the gene knockout.
Can this service be used to test combination drug strategies involving two or more agents?
Absolutely. Our service is ideally suited for testing complex dose matrices involving multiple drugs. By screening gene knockouts under the pressure of a combination therapy, we can identify targets whose deletion is synthetically lethal only in the presence of two specific compounds, providing unique insight for next-generation combination design.
What are the main advantages of using CRISPR knockout over traditional RNAi for functional screening?
CRISPR/Cas9 provides a permanent, true loss-of-function phenotype, often resulting in complete gene knockout. This is superior to RNA interference (RNAi), which provides only temporary, partial knockdown. The clean, defined knockout achieved by Creative Biolabs' service ensures that the functional consequences we observe are robust and highly reproducible.
I have a unique cancer model (e.g., a patient-derived organoid). Are your screens compatible with non-standard cell systems?
We are experts in optimizing transduction protocols for diverse and challenging cell systems, including many primary cultures and patient-derived organoids (PDOs). We assess the viability and transduction efficiency of your specific model upfront to ensure the screening delivers reliable and clinically relevant results.
Creative Biolabs' CRISPR mediated Cell Cycle Protein Knockout Screening Service is your dedicated partner in exploiting the vulnerabilities of cancer cells. By combining industry-leading CRISPR technology with deep expertise in the DNA Damage Response and cell cycle biology, we deliver high-confidence, functionally validated targets and predictive biomarkers, accelerating your path from discovery to clinical validation.
Contact Our Team for More Information and to Discuss Your Project
References
- Pellarin, Ilenia, et al. "Cyclin-dependent protein kinases and cell cycle regulation in biology and disease." Signal Transduction and Targeted Therapy 10.1 (2025): 11. https://doi.org/10.1038/s41392-024-02080-z.
- Dougherty, Michael W., et al. "Genome-scale CRISPR/Cas9 screening reveals the role of PSMD4 in colibactin-mediated cell cycle arrest." mSphere 10.3 (2025): e00692-24. https://doi.org/10.1128/msphere.00692-24.
- Distributed under Open Access license CC BY 4.0, without modification.
