CRISPR mediated Nucleoprotein Knockout Screening Service
Introduction
Nucleoproteins regulate key cellular processes like DNA repair and transcription, with novel regulators uncovered by functional genomics. Creative Biolabs' Custom CRISPR mediated Nucleoprotein Knockout Screening Service uses advanced functional genomics and high-throughput libraries to identify high-confidence nucleoprotein targets. It deciphers complex nuclear pathways, validates synthetic lethality and gene essentiality, accelerates drug discovery, and addresses nuclear factor-driven drug resistance, delivering actionable targets prioritized by functional impact in disease models.
Discover How We Can Help - Request a Consultation
CRISPR Mediated Nucleoprotein Knockout Screening Service
Background of Nucleoprotein
Nucleoproteins are proteins intimately associated with nucleic acids (DNA and RNA) and are responsible for organizing, protecting, and regulating the genetic material. This class includes transcription factors, histones, DNA and RNA polymerases, and proteins involved in DNA repair and replication—all of which are pivotal in cancer, metabolic disorders, and neurological conditions. Because they function at the intersection of genetic and signaling pathways, nucleoproteins represent high-value, yet challenging, therapeutic targets.
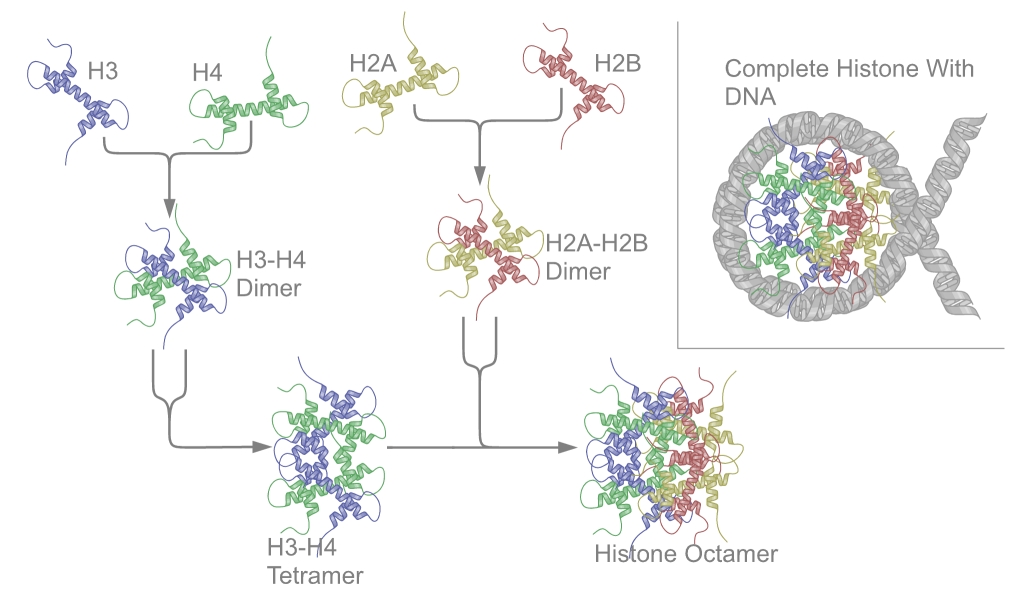 Fig.1 A schematic diagram of the assembly of core histones into nucleosomes.Distributed under CC BY-SA 3.0, from Wiki, without modification.
Fig.1 A schematic diagram of the assembly of core histones into nucleosomes.Distributed under CC BY-SA 3.0, from Wiki, without modification.
| Nucleoprotein Name/Family | Core Members | Core Functions |
|---|---|---|
| Histone Family | H2A, H2B, H3, H4 (core histones), H1 (linker histone) | Form nucleosomes (basic units of chromatin) to stabilize chromatin structure; histone modifications mediate epigenetic regulation and affect gene transcription |
| Non-histone Chromosomal Proteins | HP1 (Heterochromatin Protein 1), SMC proteins (cohesin, condensin) | HP1 maintains heterochromatin stability; SMC proteins participate in chromosome condensation and sister chromatid separation |
| Transcription-related Nucleoproteins | RNA polymerase subunits (e.g., Pol II RPB1), transcription factors (p53, NF-κB) | RNA polymerases initiate gene transcription; transcription factors regulate target gene expression and mediate signal transduction |
| Ribosomal Proteins (RP) | RPS family (e.g., RPS6), RPL family (e.g., RPL3) | Assemble with rRNA to form ribosomes and participate in protein synthesis; some regulate the cell cycle and apoptosis |
| Telomerase | hTR (RNA subunit), hTERT (reverse transcriptase), dyskerin (auxiliary protein) | Extend chromosome telomeres and maintain cell division capacity |
| Spliceosome-related Nucleoproteins | snRNP proteins (core proteins of U1, U2 snRNPs), SR proteins | Participate in pre-mRNA splicing to remove introns and ligate exons |
| DNA Repair-related Nucleoproteins | RAD51, PCNA (Proliferating Cell Nuclear Antigen) | Participate in DNA damage repair (homologous recombination, base excision repair, etc.) and maintain genome stability |
| Influenza Virus Nucleoprotein | Influenza Virus NP | Encapsidates viral ssRNA to form nucleocapsids; participates in viral replication, transcription, and particle assembly |
| SARS-CoV-2 Nucleoprotein | SARS-CoV-2 N Protein | Forms RNP complexes to stabilize the viral genome; participates in replication, packaging, and immune evasion |
| Rabies Virus Nucleoprotein | Rabies Virus N Protein | Encapsidates viral negative-strand RNA to form nucleocapsids; participates in viral replication and elicits host immune responses |
| Lamin | Lamin A/C, Lamin B | Forms nuclear lamina to maintain nuclear morphology; participates in chromatin anchoring and DNA repair |
| Ribonuclease P (RNase P) | RNA catalytic subunit, protein auxiliary subunits | Catalyzes 5' end maturation of pre-tRNA |
Tab.1 Key Nucleoproteins: Classification and Core Information.
Screening Purpose and Application
The purpose of the CRISPR mediated Nucleoprotein Knockout Screening Service is to systematically interrogate the functional consequence of losing specific nucleoprotein activity in a given cellular context. This high-throughput approach allows researchers to:
- Validate Target Essentiality: Determine if the loss of a nucleoprotein gene is lethal to a specific cancer cell line, indicating a potential therapeutic vulnerability.
- Identify Synthetic Lethality: Uncover genetic dependencies where the knockout of a nucleoprotein is only toxic when another oncogenic pathway is simultaneously compromised, facilitating the development of precise combination therapies.
- Elucidate Mechanism of Action (MoA): Define the functional role of an uncharacterized nucleoprotein in a signaling pathway, such as its influence on protein stability, as seen with p53 regulators.
The subsequent application focuses on translating these genetic findings into drug development: target prioritization, selection of lead compounds that mimic the knockout phenotype, and stratification of patients based on the specific genetic dependencies discovered.
Workflow
Creative Biolabs' robust screening process is designed for clarity, efficiency, and scalability, allowing for visualization as a comprehensive flowchart.
| Stage | Activity Description |
|---|---|
| Project Initiation & Library Design | Required Starting Materials: We require 2–3 concrete materials to initiate the project: 1) The specific nucleoprotein targets or pathway list you wish to screen (e.g., a list of 50-100 target genes); 2) Target cell line information (e.g., cell source, specific mutations like mutant p53 status); and 3) Defined functional readout (e.g., viability, protein stabilization, reporter activity). |
| High-Titer Lentiviral Production | Production of the lentiviral particles carrying the pooled or arrayed sgRNA library in our specialized clean-room facility, ensuring high quality and titer required for efficient cell transduction. |
| Genome-Wide Knockout Screening | The core high-throughput screening phase, where target cells are transduced and subjected to selective pressure (e.g., drug treatment, specific growth condition) over several passages to allow knockout phenotypes to manifest. |
| Deep Sequencing and Bioinformatics | Genomic DNA extraction from endpoint cell populations, followed by next-generation sequencing (NGS) of the integrated sgRNA sequences and advanced bioinformatics analysis using proprietary algorithms. |
| High-Confidence Target Validation | Selected top candidate genes from the initial screen are functionally validated using orthogonal methods, such as individual gene knockout, overexpression, or siRNA knockdown, confirming the primary screen findings. |
Estimated Timeframe: The typical timeframe for this service ranges from 12 to 18 weeks, depending on the complexity of the target list, the nature of the phenotypic readout, and the scope of the final validation stage.
What We Can Offer
As expert partners in functional genomics, Creative Biolabs is uniquely positioned to address the complex challenges inherent in nucleoprotein drug target discovery. We don't just run screens; we engineer success, offering a fully customizable, end-to-end service designed for maximum therapeutic impact:
Custom Library Design & Production
We deliver highly specific, tailor-made sgRNA library designs targeting your specific list of nucleoproteins or entire regulatory pathways, ensuring relevance to your precise disease model and maximum knockout efficiency.
Decisive Functional Genomics Readouts
Beyond simple gene essentiality, we specialize in advanced screening methodologies, including synthetic viability and drug-induced screens, to pinpoint critical genetic dependencies that are unique to diseased cells, drastically reducing non-actionable leads.
One-Stop Comprehensive Workflow
Creative Biolabs manages the entire process—from initial consultation and high-titer lentiviral production to advanced screening, deep sequencing, and orthogonal target validation—freeing your expert team to focus on downstream development.
Precision Targeting of Complex Regulators
We provide the expertise and tools to functionally interrogate challenging targets, such as those that regulate tumor suppressors like p53, allowing you to address the root causes of disease and resistance.
Flexible Service Scale
Whether your project requires a laboratory-scale proof-of-concept screen or a large-scale, multiplexed genome-wide campaign, our infrastructure and expert team guarantee quality and scalability across all project sizes.
Experience the Creative Biolabs Advantage - Get a Quote Today
Case Study
About half of the tumors have mutations in the p53 (TP53) gene, making it the most common mutant gene in cancer. To study the related factors regulating the stability of mutant p53, researchers conducted a screening experiment on "genes affecting the stability of mutant p53" using CRISPR gene knockout (CRISPR KO) technology. Genes such as MDM2, MDM4, USP7, HUWE1, and PPM1D/G were successfully screened out for subsequent functional verification.
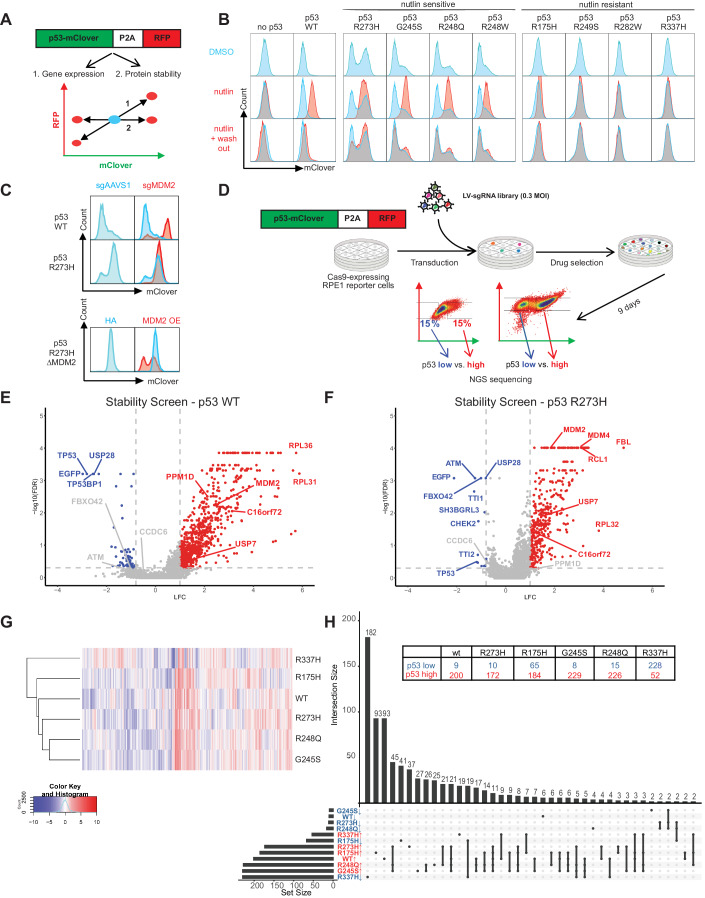 Fig.2 The regulatory factors for the stability of wild-type and mutant p53 were screened and identified by CRISPR KO technology.1
Fig.2 The regulatory factors for the stability of wild-type and mutant p53 were screened and identified by CRISPR KO technology.1
Customer Reviews

FAQs
How does the CRISPR Knockout Screen compare to traditional RNAi for target validation?
CRISPR/Cas9 achieves near-complete target protein depletion (≈100%), unlike RNAi's variable knockdown that causes ambiguous results. Our service delivers robust, high-penetrance phenotypes for definitive lead compound development.
Can the screening service accommodate my specific cell line or disease model?
Yes. Our platform is highly flexible, supporting both standard cell lines and custom screening. We optimize lentiviral transduction and selection parameters for novel or hard-to-handle client-provided models.
What's the minimum number of target genes for a custom nucleoprotein library?
We recommend at least 20 targets to ensure statistical power, with multiple sgRNAs per gene and necessary controls. We can scale up to thousands of targets based on your research needs.
What precautions ensure hit accuracy for difficult nucleoprotein targets?
We use multi-layer validation: specific sgRNAs per target, advanced bioinformatics (Bayes factors, strict FDR cutoffs), and orthogonal validation with individual knockout constructs for top candidates.
Can you help define the functional readout for targets from a genomics study?
Yes. Our expert team collaboratively designs phenotypic assays (e.g., viability, apoptosis, flow cytometry) based on your targets and disease area to maximize screening relevance.
Creative Biolabs offers the industry's most rigorous and validated CRISPR mediated Nucleoprotein Knockout Screening Service. By delivering high-confidence, actionable genetic targets and detailed mechanistic insights, we empower biopharma companies to overcome the bottleneck of target validation and accelerate the development of next-generation therapies focused on complex nuclear signaling.
Contact Our Team for More Information and to Discuss Your Project
Reference
- Lü, YiQing, et al. "Genome-wide CRISPR screens identify novel regulators of wild-type and mutant p53 stability." Molecular systems biology 20.6 (2024): 719-740. https://doi.org/10.1038/s44320-024-00032-x. Distributed under Open Access license CC BY 4.0, without modification.
