CRISPR mediated Mitochondria Knockout Screening Service
Introduction
Creative Biolabs' CRISPR mediated Mitochondria Knockout Screening Service uses precise CRISPR/Cas9 to systematically interrogate mitochondrial gene function. It generates a comprehensive functional catalog of essential mitochondrial genes, validates high-value targets, and maps gene regulation linked to OXPHOS, apoptosis, and metabolite signaling. As a trusted partner, we deliver causal data, move beyond correlative insights, and accelerate drug discovery for mitochondrial-related diseases like neurodegeneration and cancer.
Discover How We Can Help - Request a Consultation
CRISPR Mediated Mitochondria Knockout Screening Service: The Core Technology
Background of Mitochondria
Mitochondria are double-membraned organelles essential for eukaryotic life, primarily through the generation of ATP via the OXPHOS system. Beyond energy, they act as metabolic hubs, controlling fatty acid oxidation, amino acid catabolism (e.g., the BCAA pathways linked to ECHS1), iron-sulfur cluster assembly, and critical intracellular signaling cascades, notably Ca2+ signaling often mediated by the MERCS sites. Since the majority of mitochondrial proteins are encoded by the nuclear genome, these genes are accessible to powerful genomic screening tools.
Screening Purpose
The central purpose of the CRISPR mediated Mitochondria Knockout Screening Service is to overcome the limitations of traditional genetic screens by providing:
- Complete Functional Knockout: CRISPR/Cas9 ensures stable, near-100% loss-of-function, crucial for detecting phenotypes in redundant or buffered systems, unlike transient RNAi.
- Unbiased Target Identification: Systematically screen hundreds to thousands of genes against a specific mitochondrial readout to discover novel assembly factors, regulators, and non-canonical functions.
- High-Confidence Validation: Directly link a gene's loss-of-function (KO) to a measurable functional defect (e.g., loss of Complex I assembly or impaired BCAA catabolism), moving directly to actionable targets.
Subsequent Application
The output of our screening service forms the bedrock of multiple translational applications:
- Drug Target Prioritization: Use the validated gene list to initiate HTS campaigns for small molecule modulators (activators or inhibitors) of the identified pathways (e.g., targeting RTN4IP1 to rescue CI function).
- Biomarker Discovery: Identify genes whose expression level correlates with the functional defect, serving as predictive or prognostic biomarkers for mitochondrial diseases.
- Mechanism of Action Studies: Employ the single-gene KO lines for in-depth mechanistic studies to understand how lead compounds exert their effects on mitochondrial structure and function.
Workflow
The process is designed for maximum clarity, precision, and reproducibility, offering a clear roadmap for clients.
Required Starting Materials
To initiate our specialized screening service, clients typically provide:
- List of Target Genes: A prioritized list of 100-1,500 nuclear-encoded genes hypothesized to regulate mitochondrial function (e.g., genes identified from proteomics or GWAS studies).
- Disease Cell Line: Specific, well-characterized cell lines (e.g., patient-derived iPSCs, cancer lines, or immortalized primary cells) relevant to the target disease state.
- Phenotypic Assay Parameters: Detailed protocols or specifications for the readout of interest (e.g., acceptable signal-to-noise ratio for the oxygen consumption rate (OCR) assay).
| Step | Description of Activities |
|---|---|
| Library Design & Construction | Design highly active single-guide RNAs (sgRNAs) targeting every gene in the client's custom list, along with non-targeting controls. sgRNAs are synthesized and cloned into lentiviral vectors. |
| Cell Transduction & Selection | Transduce the client's disease-relevant cell line with the pooled lentiviral library at a low Multiplicity of Infection (MOI) to ensure single-sgRNA integration per cell. Select for transduced cells using an antibiotic resistance marker. |
| Phenotypic Screening | Subject the pooled cells to the selection pressure (e.g., galactose media for OXPHOS dependency, specific metabolite concentrations, or a neurotoxic challenge). This includes time-resolved data collection and high-content imaging analysis. |
| Deep Sequencing & Hit Calling | Extract genomic DNA from cells before and after the phenotypic selection. Amplify the sgRNA region and quantify the enrichment or depletion of specific sgRNAs using Next-Generation Sequencing (NGS). |
| Validation & Mechanism Elucidation | Construct individual knockout cell lines for the top hits (e.g., RTN4IP1, GET4). Validate the phenotype using orthogonal assays (e.g., Seahorse analysis, complexome profiling, Ca2+ imaging). |
Final Deliverables
Upon completion, clients receive specific, high-value data packages:
- Annotated Gene Catalog: A list of validated hit genes, ranked by phenotypic severity (e.g., Z-scores), ready for downstream drug screening.
- Raw and Processed NGS Data: All sequencing files and computational analysis reports for full transparency and in-house data mining.
- Mechanism of Action Report: Detailed functional data (e.g., Western blots, Seahorse plots, complexome heatmaps) confirming the mechanism of the top 5 validated hits.
Estimated Timeframe
The typical timeframe for this comprehensive service ranges from 12 to 18 weeks, depending on the complexity of the client's cell model and the stringency of the required phenotypic assay development.
What We Can Offer
At Creative Biolabs, we understand that every research project is unique, requiring specialized tools and flexible service execution. Our CRISPR mediated Mitochondria Knockout Screening Service is not a one-size-fits-all solution; it is a fully customizable, end-to-end platform designed to fit the precise mechanistic questions and scale of your drug discovery program.
Custom Library Design & Construction
We offer the flexibility to design focused sgRNA libraries against any subset of the nuclear-encoded mitochondrial genome, ensuring high efficiency and low noise tailored to your specific disease pathway or target hypothesis (e.g., exclusively targeting OXPHOS assembly factors or MERCS regulators).
One-Stop Functional Screening Service
We provide integrated service from initial target gene selection, custom library design, stable cell line generation (including challenging iPSC or patient-derived models), through to large-scale, high-content phenotypic screening.
High-Resolution Phenotypic Readouts
We run screening assays in batch or continuous mode using advanced functional readouts—including time-resolved Oxygen Consumption Rate (OCR) and Ca2+ flux—going beyond simple viability to provide high-fidelity, functional data.
Customized Mechanistic Validation
The core service includes the option for deep validation using technologies like Complexome Profiling (to track assembly intermediates, as required for RTN4IP1 analysis) or targeted Metabolomics (to validate non-canonical pathways, as seen with ECHS1).
GMP-Ready Data Packages
All data and documentation procedures adhere to well-established quality systems, ensuring the necessary rigor for targets intended for clinical translation, supporting your Quality-by-Design (QbD) requirements.
Guaranteed Stability and Quality
We guarantee the stability and viability of the cell banks used throughout the process and utilize high-standard Quality Control (QC) tools to quantify and evaluate the quality of the final validated hits.
Dedicated Scientific Consultation
We provide ongoing expertise to optimize culture and screening conditions to maximize phenotypic yield and translate raw data into actionable therapeutic strategies, ensuring successful downstream applications.
Experience the Creative Biolabs Advantage - Get a Quote Today
Case Study
OXPHOS-deficient cells can survive in a culture medium rich in glucose, but they cannot survive when glucose is replaced by galactose. However, traditional whole-genome screening based on galactose, if conducted only at early time points, often fails to identify the key regulatory components of mitochondrial oxidative metabolism. To overcome this inherent limitation, we have constructed a novel lentiviral library covering all the nuclear-coding mitochondrial genes of humans. After screening, we collected cells at the initial time point (t0h) and multiple subsequent time points (t24h, t48h, 1 week, 2 weeks), and isolated genomic DNA (gDNA) for sequencing. Two genes with unclear characteristics - RTN4IP1 and ECHS1 - were identified as results, and they showed significant correlations with the known OXPHOS components.
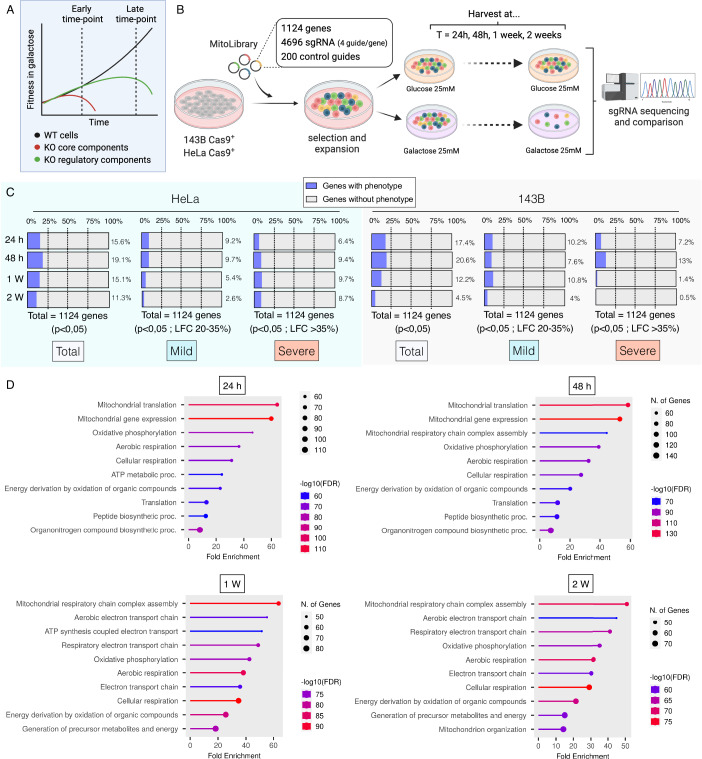 Fig.1 Use the custom-developed Mito library for galactogen-based CRISPR screening.1
Fig.1 Use the custom-developed Mito library for galactogen-based CRISPR screening.1
Customer Reviews

FAQs
How does your targeted CRISPR Mito-library compare to a full genome-wide screen for mitochondrial disease research?
A full genome-wide screen offers breadth but often lacks the statistical power to resolve subtle mitochondrial regulators. Our targeted Mito-library contains only the nuclear-encoded mitochondrial genes, leading to deeper coverage, significantly less noise, and increased sensitivity. This precision is why we can identify complex targets like GET4/MERCS regulators and Complex I assembly factors (RTN4IP1) that are missed in general screens. If you have a preliminary target list, our focused approach provides unparalleled efficiency.
What is the most critical technical challenge you address, and how do you ensure the functional relevance of the hit genes?
The primary challenge is linking a gene KO to a functional cellular defect. We address this by focusing on sophisticated, quantitative readouts like the Oxygen Consumption Rate (OCR) using Seahorse technology and advanced high-content imaging of Ca2+ dynamics. This moves beyond simple cell viability to ensure your hits are true modulators of mitochondrial physiology, which is crucial for therapeutic success.
Can your service be adapted for my specific disease model, such as patient-derived iPSCs or a unique cancer type?
Absolutely. Our service is designed for maximum flexibility. The first step involves rigorous optimization of the lentiviral transduction and phenotypic assay conditions for your specific cell line. We have extensive experience with challenging models, including terminally differentiated cells and primary cells. Let's discuss your specific model in detail to tailor the perfect strategy.
How do you handle the potential off-target effects that are sometimes associated with CRISPR technology?
We employ a multi-layered mitigation strategy. We use best-in-class sgRNA design algorithms to minimize off-target risk and include multiple independent sgRNAs per gene to ensure that only consensus hits are carried forward. Furthermore, our final step includes orthogonal validation in single-clone KO lines, providing definitive confirmation that the observed phenotype is due solely to the loss of the target gene.
What level of support is offered after the final data and reports are delivered?
Our support does not end with the final report. As expert biology specialists, we offer post-delivery scientific consultation to assist your team with data interpretation, follow-up mechanism-of-action studies, and strategic planning for downstream drug development efforts. Our goal is to ensure you effectively translate the screening data into clinical leads.
Creative Biolabs is committed to being the industry leader in precision mitochondrial biology. Our CRISPR mediated Mitochondria Knockout Screening Service provides the high-confidence, functionally validated targets required to tackle complex metabolic and neurodegenerative diseases. Partner with us to unlock the next generation of mitochondrial therapeutics.
Contact Our Team for More Information and to Discuss Your Project
Reference
- Zamora-Dorta, Marcos, et al. "Time-resolved mitochondrial screen identifies regulatory components of oxidative metabolism." EMBO reports 26.12 (2025): 3045-3074. https://doi.org/10.1038/s44319-025-00459-9. Distributed under Open Access license CC BY 4.0, without modification.
