CRISPR mediated Membrane Protein Activation Screening Service
Introduction
Functional genomics bridges 'omics' data and therapeutic targets, and our CRISPR-mediated Membrane Protein Activation Screening Service leads this field. Leveraging CRISPRa to systematically activate genes, it identifies causal modulators of disease phenotypes, aiding in discovering metastasis drivers, host-virus factors, and engineering cell lines. Focusing on membrane proteins and non-coding regions, Creative Biolabs delivers definitive functional validation via proprietary high-throughput CRISPRa screening and advanced 3D biomimetic platforms. We turn genomic association into engineered cellular function, minimizing discovery risk and accelerating actionable drug leads.
Discover How We Can Help - Request a Consultation Today
CRISPR Mediated Membrane Protein Activation Screening Service
Membrane proteins—including ion channels, GPCRs, and transporters—represent approximately 60% of all current drug targets due to their critical roles in cellular communication and environmental sensing. However, their complex structure and low expression levels make them notoriously difficult to study, purify, and functionally validate in traditional screening formats.
Screening Purpose
The core purpose of our CRISPR mediated Membrane Protein Activation Screening Service is to bypass the limitations of transient transfection and recombinant expression. By using the CRISPRa system (which recruits transcriptional activators to endogenous promoter regions), we systematically overexpress or activate native membrane proteins in situ. This allows us to rapidly determine which proteins, upon upregulation, drive a specific desired phenotype, such as:
- Enhanced T-cell activity in an immune checkpoint assay.
- Increased resistance or sensitivity to a compound in a drug screen.
- Significantly higher protein folding and expression yields in bioproduction lines.
Subsequent Application
Hits identified through this activation screening service are immediately validated, providing highly specific, druggable targets.
| Application Field | Specific Directions | Detailed Description |
|---|---|---|
| In-depth Analysis of Disease Mechanisms | Protein Function Validation | Confirm its core role in cell proliferation, invasion, signal transduction, or pathogen infection (e.g., whether it is a key regulator of tumor metastasis or a viral entry receptor) through overexpression, knockdown, and other experiments. |
| Molecular Pathway Exploration | Identify upstream and downstream interacting partners of the protein (e.g., binding receptors, signaling molecules), map the signaling network it participates in, and reveal novel mechanisms underlying disease occurrence and progression. | |
| Drug Target Development and Screening | As a Drug Target | Design small-molecule inhibitors, neutralizing antibodies, or peptide drugs targeting the natural conformation of the membrane protein to block its abnormal activity (e.g., development of inhibitors for oncogenic membrane proteins in tumors). |
| High-Throughput Drug Screening | Use the target membrane protein as a screening target to construct cell models or in vitro binding systems, and rapidly screen potential drugs that can regulate its function from compound libraries. | |
| Diagnostic Biomarkers and Prognostic Evaluation | Development of Diagnostic Tools | Detect the expression level of the target membrane protein in patients' tissues and blood to verify its potential as a biomarker for early disease diagnosis and classification (e.g., immunohistochemical detection reagents for tumor-specific membrane proteins). |
| Prognostic Prediction | Analyze the correlation between protein expression levels and patients' survival time and treatment response, construct prognostic evaluation models, and guide the selection of clinical treatment regimens. | |
| Cell Engineering and Therapeutic Optimization | Cell Therapy Target Modification | Introduce screened functional membrane proteins (e.g., chemokine receptors) into engineered cells such as CAR-T to enhance their tumor-targeting infiltration ability or killing efficiency. |
| Oncolytic Virus/Vaccine Optimization | Use the membrane protein as a target to modify the surface ligands of oncolytic viruses for enhanced targeted infection of tumor cells, or design vaccines against this protein to activate the body's specific immune response. |
Workflow
Required Starting Materials
- Target Gene IDs (with provided cDNA or gDNA sequences), serving as the core targeting basis for CRISPRa screening;
- Customer-specified cell lines (e.g., HEK293T, CHO cells, or primary cells), which need to be compatible with the requirements of subsequent screening experiments;
- Target phenotype assay methods (e.g., viral titer detection, T-cell killing efficiency measurement, protein yield analysis, etc.), used to define the functional evaluation criteria in the screening process.


Library Design & Construction
Custom design of CRISPRa guide RNA (sgRNA) libraries for specific promoter regions or whole genomes to ensure targeting accuracy;
Cell Line Engineering & Quality Control
Construction of stable and efficient cell lines using Cas9 fusion proteins (e.g., dCas9-activator), followed by comprehensive quality control to verify the applicability of the cell lines;


High-Throughput Functional Screen
Conduct large-scale screening under strictly controlled experimental conditions, with commonly used techniques including flow cytometry (FACS), high-content imaging, or deep sequencing screening;
Hit Selection & Data Deconvolution
Deconvolute complex screening data through bioinformatics analysis, and prioritize high-confidence candidate targets based on statistical reliability and pathway relevance;


Secondary Validation
Independently validate core candidate targets in targeted experiments, including dose-response curve determination and alternative functional tests, to confirm the functional stability of the targets.
Final Deliverables
- Validated Target List: Includes functionally confirmed genes, ranked by the degree of phenotypic impact and therapeutic potential, providing priority references for subsequent research;
- Next-Generation Sequencing (NGS) Data & Raw Reads: Provides complete and annotated NGS data and raw output results to ensure data transparency and support customers' in-house validation;
- Mechanistic Report: Details the process of pathway deconvolution and secondary validation data, while providing strategic recommendations for downstream drug development to facilitate the implementation of subsequent research.


Estimated Timeframe
The typical timeframe for this comprehensive service ranges from 10 to 14 weeks, depending on the complexity of the initial gRNA library design and the biological complexity of the target phenotype assay.
What We Can Offer
Bespoke Library Design
Customized CRISPRa gRNA library construction targeting specific gene sets, protein domains, or entire genomes to perfectly align with your research focus (e.g., specific receptor families, non-coding regions, or disease pathways).
Physiologically Relevant Models
Precision screening integration into 3D multicellular tumor spheroids, co-culture systems, or species-specific cell lines to ensure functional validation is grounded in a clinically or biologically accurate context.
Gain-of-Function Phenotyping
Systematic functional analysis using CRISPRa to identify low-abundance membrane proteins or silent therapeutic targets that, when upregulated, causally drive a desired phenotype.
Production Cell Line Optimization
Targeted CRISPR editing services to engineer high-performance cell lines, significantly enhancing the expression and functional folding of notoriously challenging biologics, such as therapeutic membrane proteins.
Customizable Workflow & Scale
Flexible project design ranging from focused screens on a few hundred targets to large-scale genome-wide screens, ensuring the service is precisely scaled to meet your project timeline and budget requirements.
Experience the Creative Biolabs Advantage - Get a Quote Today
Case Study
To screen for tumor cell surface proteins that may inhibit the functions of NK cells and CD8+ T cells, a study was conducted using functional-acquired screening using the CRISPRa library. The results showed that MUC21 was the most important target in the NK cell escape mechanism. After screening out this target, further in vivo or in vitro experiments can be conducted to detect its mechanism of action. MUC21 inhibits the killing function of NK cells, reduces the secretion of CD107a and IFN-γ, and the expression of MUC21 on the surface of cancer cells also blocks the interaction between cancer cells and immune cells.
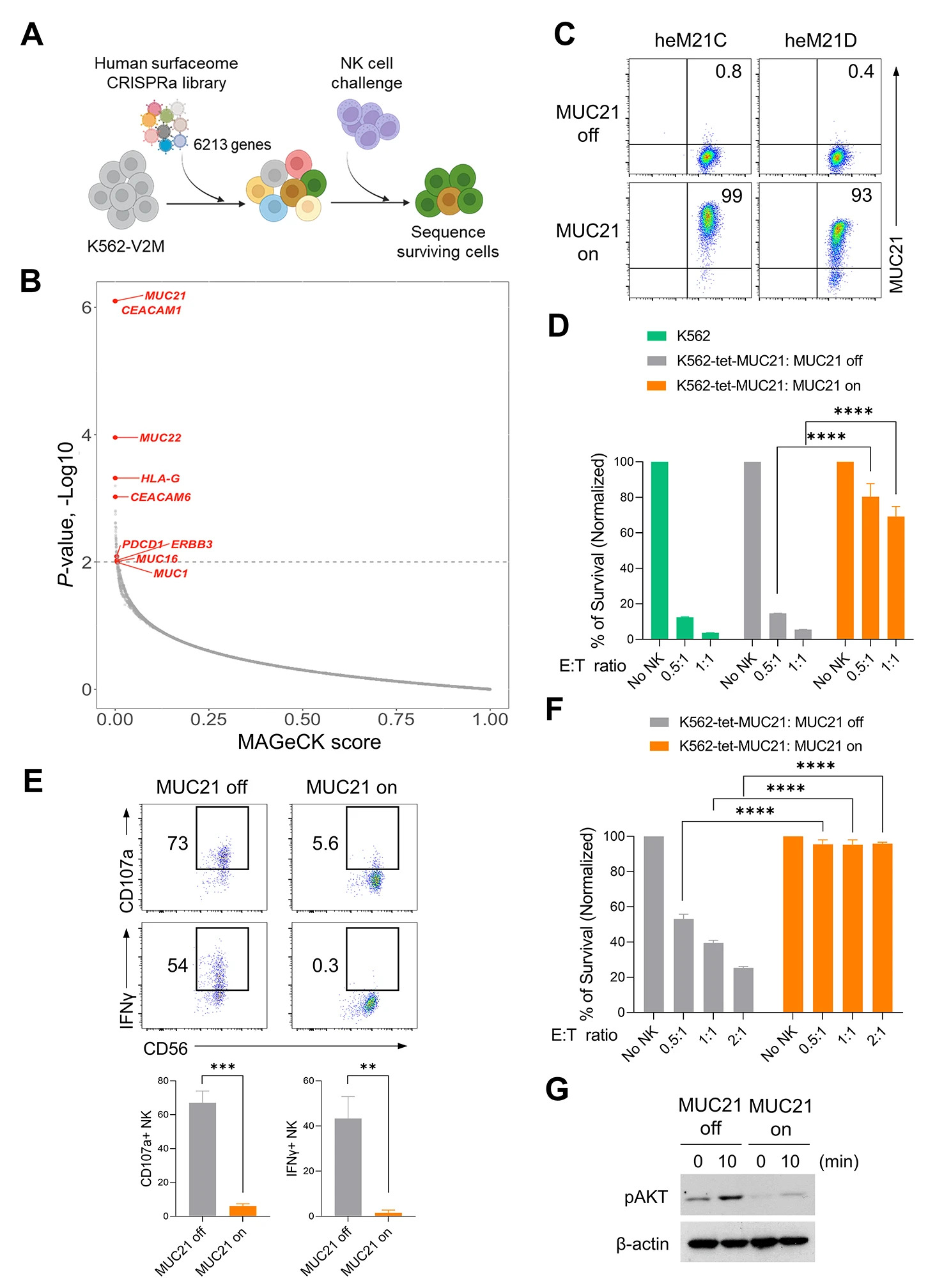 Fig.1 The CRISPRa screening technology was utilized to identify genes that might inhibit the toxicity of NK cells.1
Fig.1 The CRISPRa screening technology was utilized to identify genes that might inhibit the toxicity of NK cells.1
Customer Reviews

FAQs
How does CRISPRa activation screening compare to traditional CRISPR KO (knockout) screening for target validation?
CRISPR KO screens identify genes necessary for a function (loss-of-function), while our CRISPRa activation screening identifies genes sufficient to drive a function (gain-of-function). For many therapeutic targets—especially receptors or low-abundance regulatory proteins—activation provides a more direct and physiologically relevant model of potential drug action.
Can your service be utilized with complex 3D or patient-derived cell models?
Absolutely. Our protocols are optimized for integration into advanced systems, including 3D tumor spheroids and host-virus interaction models. Screening in these physiologically relevant environments ensures that the targets identified are functional within the complex cell-cell and cell-matrix interactions that mirror the in vivo environment, boosting your confidence in translation.
What measures do you take to minimize off-target effects and false positives in the activation screen?
Creative Biolabs employs a multi-pronged quality control strategy. We utilize highly optimized and published guide RNA libraries with validated specificity, typically targeting multiple sites per gene. Furthermore, our final validation stage utilizes secondary, orthogonal assays to confirm that the observed phenotype is truly mediated by the target gene's upregulation, resulting in high-confidence data.
Is this service solely focused on cancer or membrane proteins?
While we specialize in CRISPR-mediated Membrane Protein Activation Screening, the underlying functional genomics platform is highly versatile. We successfully apply similar principles to infectious diseases (host factor screens), metabolic disorders, and complex cell line engineering for bioproduction. We encourage you to detail your unique project for a customized approach.
Creative Biolabs' CRISPR mediated Membrane Protein Activation Screening Service delivers the future of drug discovery today: functional certainty, reduced attrition, and accelerated timelines. Leveraging over 20 years of scientific leadership and proprietary high-throughput platforms, we transform challenging genomic data into validated, high-value therapeutic targets across oncology, infectious disease, and bioproduction.
Contact Our Team for More Information and to Discuss Your Project
Reference
- Lee, Dong-hee, et al. "A CRISPR activation screen identifies MUC-21 as critical for resistance to NK and T cell-mediated cytotoxicity." Journal of Experimental & Clinical Cancer Research 42.1 (2023): 272. https://doi.org/10.1186/s13046-023-02840-9. Distributed under Open Access license CC BY 4.0, without modification.
