CRISPR mediated Membrane Protein Inhibition Screening Service
Introduction
Creative Biolabs' CRISPR mediated Membrane Protein Inhibition Screening Service uses functional genomics precision to identify/validate novel membrane protein drug targets (e.g., GPCRs, SLC transporters) critical in cancer, infectious resistance, and other diseases. It employs context-aware screening tailored to complex disease microenvironments like TME or antibiotic stress. The service accelerates target discovery, quantifies membrane proteins' functional necessity, addresses therapeutic resistance and off-target toxicity, and specializes in disease-specific essential targets to maximize therapeutic specificity.
Discover How We Can Help - Request a Consultation
CRISPR Mediated Membrane Protein Inhibition Screening Service
Background of CRISPR Inhibition
We employ CRISPR Interference (CRISPRi), a powerful method for reversible and tunable gene suppression. The system uses a catalytically inactive Cas9 (dCas9) fused to a transcriptional repressor. This complex is guided by a specific single-guide RNA (sgRNA) to the promoter region of a target gene (e.g., a membrane protein). By physically blocking RNA polymerase access, the system achieves dose-responsive knockdown of gene expression. This offers a superior alternative to traditional gene knockout, as it avoids genetic drift and allows the study of genes that are otherwise essential for basic cell survival.
Screening Purpose
The primary purpose is to perform high-throughput, loss-of-function screens across the entire set of membrane protein-encoding genes (or a focused subset). This systematically reveals which membrane protein's inhibition or suppression leads to a desired phenotypic outcome, such as:
- Increased sensitivity to an existing therapeutic agent (drug synergy).
- Growth impairment under environmental stress (e.g., TME-induced nutrient starvation).
- Bacterial cell death or reduced virulence.
Subsequent Application
The outputs from the inhibition screening drive several critical downstream applications:
- Drug Compound Testing: Validated targets are immediately leveraged for screening small molecule libraries to find compounds that mimic the CRISPRi-induced functional inhibition.
- Mechanism of Action (MoA) Studies: Hits are prioritized for detailed biological investigation, helping clients understand the exact cellular machinery disrupted by inhibiting the membrane protein (e.g., how serotonin uptake is linked to ferroptosis resistance).
- Clinical Candidate Prioritization: Targets proven essential under in vivo-mimicking conditions are moved forward with high confidence, de-risking the entire preclinical pipeline.
Workflow
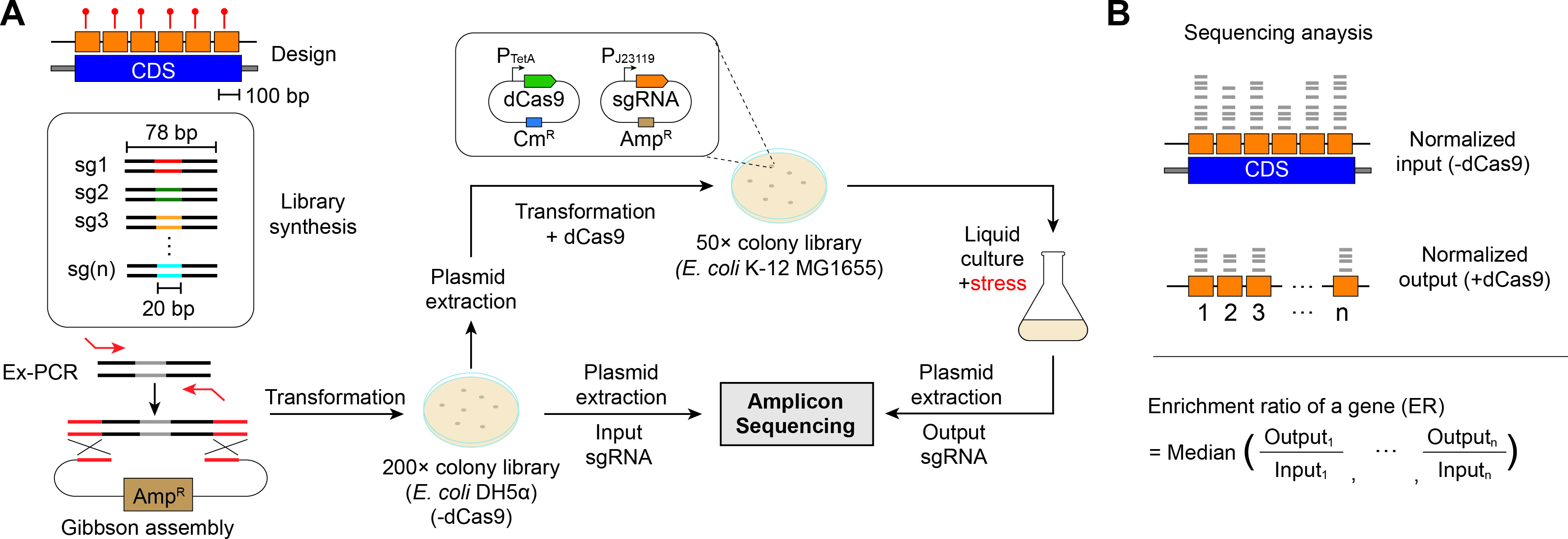 Fig.1 Genome-wide gene adaptability analysis of Escherichia coli was conducted using CRISPRi library screening.1,2
Fig.1 Genome-wide gene adaptability analysis of Escherichia coli was conducted using CRISPRi library screening.1,2
The typical timeframe for this service ranges from 10 to 14 weeks, depending on the complexity of the required cell line engineering and the chosen screening environment (e.g., TME-mimicking conditions).
Required Starting Materials
To initiate a project, clients typically provide:
- Target Gene List: A prioritized list of genes, such as the entire SLC transporter or GPCR family, for focused screening.
- Cell Lines: Client-provided or custom-sourced disease models (e.g., PDAC cell lines for oncology or bacterial strains like E. coli for AMR).
- Assay Readout Criteria: Clear definitions for the desired phenotypic endpoint (e.g., cell viability under nutrient starvation, resistance to a specific drug, or pathway activation).


Target Definition and sgRNA Library Design
Collaborate to define screening scope (genome-wide/focused) and design high-resolution sgRNA libraries, guiding dCas9 for optimal, tunable target gene inhibition.
Cell Line Engineering and Transduction
The Engineer selected cell lines to stably express CRISPRi machinery (dCas9), then transduced pooled sgRNA libraries via high-efficiency lentivirus for deep target coverage.


Phenotypic Screening in Context
Culture cells under physiologically relevant stress (e.g., TME low-glucose, anaerobic antibiotic conditions); enrich/deplete cells based on whether inhibited genes aid or harm cell survival.
Next-Generation Sequencing and Data Analysis
Isolate genomic DNA from initial/final cell pools, amplify sgRNA cassettes for NGS, and calculate Enrichment Ratios (ER) to quantify each gene's fitness effect.


Hit Validation and Prioritization
Validate top hits with single-gene sgRNAs to confirm effects, and provide mechanistic hypotheses for pathway involvement.
Final Deliverables
on project completion, you will receive:
- Gene Essentiality Heatmap: A high-resolution graphical representation comparing gene fitness across the tested environments (e.g., TME vs. standard media).
- Prioritized Hit List: A quantitative list of validated, context-dependent targets (e.g., membrane transporters or regulatory components like CpxR).
- Mechanism of Action (MoA) Hypotheses Report: Detailed biological interpretations supporting the functional role of the prioritized hits, linking them to specific disease survival mechanisms.

What We Can Offer
Customized Library Design
Full flexibility in designing sgRNA libraries—from genome-wide screens to focused libraries targeting specific membrane protein families (e.g., SLC, GPCR, Ion Channels) to match your therapeutic area.
Tunable Target Suppression (CRISPRi)
Utilize CRISPR Interference (CRISPRi) for stable, dose-responsive gene knockdown that accurately models small molecule inhibition, avoiding the complexities of complete genetic knockout.
Physiologically Relevant Contexts
Screening is performed in custom-designed, stress-inducing microenvironments, including TME-mimicking low-nutrient media and anaerobic infection models, guaranteeing the identification of targets that matter in vivo.
Direct Pathway Dissection
The platform is engineered to uncover and validate entire regulatory nodes (like the CpxR system in AMR) and novel, critical survival pathways (like the serotonin/ferroptosis link) for re-sensitization strategies and unique drug mechanisms.
High-Resolution Quantitative Data
Delivery of clear, actionable data, including Enrichment Ratios (ER) and comprehensive Mechanism of Action (MoA) Hypotheses Reports for immediate pipeline integration.
Experience the Creative Biolabs Advantage - Get a Quote Today
Customer Reviews

FAQs
How does CRISPRi screening compare to traditional siRNA or shRNA screening for membrane proteins?
CRISPRi offers superior specificity and knockdown uniformity compared to transient siRNA transfection or shRNA. The stable expression of dCas9 results in more consistent and tunable inhibition, leading to more reproducible and quantitative data, essential for reliable Gene Essentiality (ER) calculations.
Can your platform identify targets that prevent antibiotic resistance?
Absolutely. By screening under sub-lethal antibiotic concentrations and harsh environmental conditions (e.g., low oxygen), we identify genes that, when inhibited, re-sensitize the bacteria. This led to the discovery of regulatory nodes like the CpxR system, which controls membrane depolarization and drug uptake.
Is your screening platform effective for membrane targets beyond cancer, such as GPCRs or ion channels?
Yes. While we highlight TME and AMR, the core platform applies to any membrane protein class. By utilizing a functional sgRNA library targeting these families and pairing it with a relevant physiological assay (e.g., calcium flux or signaling pathway activation), we can pinpoint and validate functional regulatory members in any disease area.
How do you ensure the targets identified in TME-mimicking conditions are clinically relevant?
We prioritize Contextual MoA by designing the screening environment to strictly mirror the nutrient and oxygen deprivation found in vivo. Our successful identification of TME-essential nutrient transporters demonstrates that our platform finds targets that are selectively required for tumor survival, minimizing the risk of systemic toxicity in a clinical setting.
What is the benefit of generating an Enrichment Ratio (ER) rather than just a survival score?
The ER provides a quantitative, high-resolution measure of fitness effect, which is crucial for understanding essential genes. By showing the relative degree of fitness change upon inhibition, we can distinguish between highly valuable, essential targets and those with only marginal effects, accelerating the focus on high-impact therapeutic leads.
Creative Biolabs' CRISPR mediated Membrane Protein Inhibition Screening Service provides the critical link between complex genomic data and actionable drug targets. By offering single-cell resolution and context-aware functional screening, we enable our clients to discover and validate targets across oncology, infectious disease, and neuroscience with unprecedented confidence.
Contact Our Team for More Information and to Discuss Your Project
References
- Choe, Donghui, et al. "CRISPRi screening reveals E. coli's anaerobic-like respiratory adaptations to gentamicin: membrane depolarization by CpxR." mSystems (2025): e00353-25. https://doi.org/10.1128/msystems.00353-25.
- Distributed under Open Access license CC BY 4.0, without modification.
