CRISPR mediated Cancer related Gene Knockout Screening Service
Introduction
Service uses advanced high-throughput functional genomics to systematically interrogate cancer genomes. It uncovers therapeutic vulnerabilities by linking gene loss-of-function to cell death, delivering causal dependency data. As experts in functional genomics, we accelerate target discovery, de-risk preclinical assets, and solve oncology pipeline challenges, enabling next-generation, precise therapeutics.
CRISPR Mediated Cancer-Related Gene Knockout Screening Service
Background of Cancer-Related Genes
Cancer progression is driven not only by primary mutations but also by epigenetic modifications and the surrounding supportive stroma. Epigenetic regulators are frequently overexpressed in malignancies. These factors dynamically control the stability and translation of thousands of transcripts, essentially acting as master switches for pro-oncogenic programs like proliferation and metastasis. Similarly, elements of the TME, such as Cancer-Associated Fibroblasts (CAFs), activate resistance pathways, demanding targets that disrupt this protective shield. Identifying these non-mutational vulnerabilities is where CRISPR screening offers a distinct advantage.
Screening Purpose
The primary purpose of our CRISPR knockout screening service is to determine the functional necessity of cancer-related genes. While gene expression data can provide hints, only loss-of-function screening can definitively prove that a cancer cell is dependent on a specific gene for survival.
Our service aims to:
- Identify Core Dependencies: Pinpoint essential genes (e.g., metabolic enzymes, signaling mediators, transcription factors) required for cell viability.
- Validate Novel Targets: Provide functional evidence to confirm candidates identified through other methods (e.g., proteomics, genomics).
- Uncover Resistance Mechanisms: Screen for genes whose knockout resensitizes cells to existing therapies, revealing crucial pathways for combination strategies.
Subsequent Application
The actionable data generated by Creative Biolabs' CRISPR screening platform directly informs critical downstream applications:
- Lead Target Selection: High-confidence hits (genes with low tolerance for knockout) become the highest-priority targets for small-molecule inhibitor or biologic development.
- Biomarker Discovery: Genes that show dependency only in a specific cancer subtype or in a certain mutation (e.g., MYCN amplification) can be developed into highly specific companion diagnostic biomarkers.
- Combination Therapy Rationalization: Identifying synergistic or synthetic lethal interactions provides the intellectual property foundation for next-generation drug combinations, moving assets rapidly into preclinical testing.
Workflow
Required Starting Materials
- Cell Line/Model: Specific cancer cell line(s) (e.g., patient-derived organoids (PDOs), 2D/3D culture) and corresponding genetic information.
- Target List: A focused set of genes (e.g., epigenetic regulators like KIAA1429, TME-related factors) or a genome-wide library for comprehensive screening.
- Research Goals: Clearly defined phenotypic readout (e.g., survival in specific drug concentrations, proliferation rate, metastatic potential).


Library Design & Synthesis
Custom design of high-quality single guide RNA (sgRNA) libraries, ensuring optimal coverage and minimizing off-target effects.
Cell Line Engineering
Stable integration of Cas9 into the client's chosen cancer model (e.g., BE(2)-C Neuroblastoma cells) to establish a robust screening platform.


Pooled Screening & Selection
High-throughput transduction of the sgRNA library followed by selective pressure (drug treatment, nutrient deprivation, specific growth conditions) to enrich for surviving cells or deplete dying cells.
Deep Sequencing & Bioinformatics Analysis
Extraction of genomic DNA and deep sequencing of integrated sgRNAs. Advanced computational methods (including robust statistical models) are applied to compare guide abundance before and after selection.


Validation & Hit Prioritization
Top candidate genes undergo secondary validation (e.g., individual gene knockout, dose-response studies) to confirm their dependency and prioritize the most promising drug targets.
Final Deliverables
- Raw & Processed Sequencing Data: Comprehensive data files detailing sgRNA counts for every time point and condition.
- Validated Target Report: A ranked list of essential genes with calculated Gene Dependency Scores and supporting visual data, clearly illustrating the causal role of each hit.
- Mechanistic Insights: Pathway enrichment analysis and preliminary interpretation linking the identified genes to known cancer hallmarks (e.g., m⁶A regulation, TME signaling).


Estimated Timeframe
The typical timeframe for a focused screening project ranges from 10 to 16 weeks, depending on the complexity of the cell model (2D vs. 3D organoids) and the size of the gene library being investigated.
Discover How We Can Help - Request a Consultation
What We Can Offer
As an excellent seller and your dedicated partner in functional oncology, Creative Biolabs is committed to transforming your target hypotheses into validated clinical candidates. We understand that success hinges on precision and predictability, which is why our Custom CRISPR Mediated Gene Knockout Screening Service is engineered for maximum translational value.
Full Customization for Unique Research Goals
We provide a one-stop functional genomics solution that is entirely tailored to your specific cancer model and research question, whether you require a focused screen on epigenetic drivers or a complex drug resistance screen.
Precision Library Design
Our expertise ensures the custom design of high-quality single guide RNA (sgRNA) libraries optimized for your specific targets to maximize on-target editing efficiency.
High-Throughput Screening in Predictive Models
Leverage our capacity for high-throughput execution in advanced, clinically relevant systems, including patient-derived organoids (PDOs) and TME co-culture assays, providing data that accurately reflects in vivo disease complexity.
Rigorous Quality-by-Design (QbD) Workflow
Implementation of strict quality control (QC) protocols throughout the process—from Cas9 integration to deep sequencing—guaranteeing the highest standard of data integrity and reproducibility.
Actionable Bioinformatics and Mechanistic Insights
We go beyond raw counts, applying advanced computational methods and Gene Dependency Scoring to convert complex data into a clear, prioritized list of functionally validated targets for immediate downstream drug development.
Flexible Capacity and Scalability
Creative Biolabs offers flexible capacity to handle all project scales, from small pilot screens to validate a preliminary target list to large-scale, multiplexed functional screens, ensuring rapid and efficient resource allocation.
Case Studies
To systematically analyze tumor-dependent genes in Ewing's sarcoma (ES), researchers used CRISPR technology to perform in vitro screening of epigenetic/transcriptional regulators in two commonly used ES cell lines (A673, SKNMC), identifying 369 and 297 tumor-dependent genes, respectively. Further in vivo CRISPR knockout screening identified 319 and 85 tumor-dependent genes in the two cell lines. Comprehensive analysis revealed 42 ES-dependent epigenetic/transcriptional regulators, including known ones like AURKB, CDK9, MYC, RPA2, SSRP1, and SUPT16H. Functional enrichment analysis showed RNA processing was the most highly enriched function.
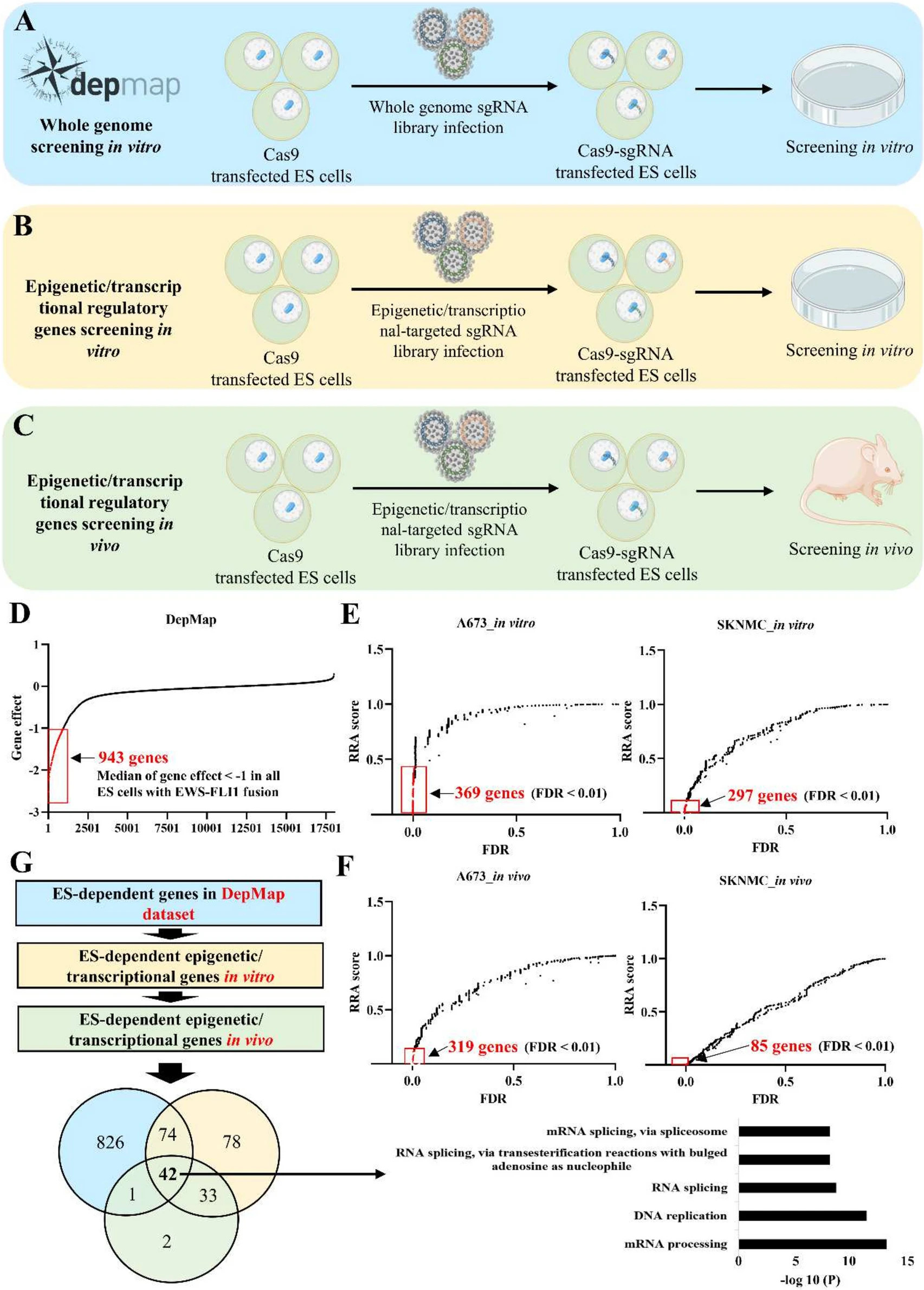 Fig.1 CRISPR-Cas9 gene knockout screening for the identification of ES-dependent genes.1
Fig.1 CRISPR-Cas9 gene knockout screening for the identification of ES-dependent genes.1
Customer Reviews

Experience the Creative Biolabs Advantage - Get a Quote Today
FAQs
How does Creative Biolabs' screening compare to standard genetic screens?
Our platform focuses on generating causal, functional data rather than just correlation. We specialize in complex, custom models (like PDOs and co-culture systems) to capture TME relevance, providing a far more predictive result set for target validation than standard cell-line screens.
What level of sequencing depth do you provide, and is the data easy to interpret?
We utilize deep sequencing to ensure high coverage and robust statistical power, even for low-abundance guides. The final deliverable includes a high-confidence, ranked list of hits with clear Gene Dependency Scores, allowing immediate prioritization without the need for extensive internal bioinformatics resources.
Can this service help identify targets related to non-coding RNA or epigenetic mechanisms like m6A?
Absolutely. Our custom and focused libraries are often designed specifically to interrogate epigenetic regulators (writers, readers, and erasers, like KIAA1429) and non-coding RNA processing factors, which are often highly selective vulnerabilities in cancer.
What is the typical turnaround time for a genome-wide screen versus a focused library screen?
Genome-wide screening often requires 14–18 weeks due to the volume of sequencing, while a focused library screen (e.g., targeting a specific pathway) typically ranges from 10–14 weeks. Project duration is confirmed upon consultation based on model complexity.
We have a preliminary target list from RNA-Seq. Can your service help de-risk these candidates quickly?
Yes, this is our core strength. We can rapidly transition your preliminary candidates into a focused, low-cost CRISPR validation screen. This immediately provides the functional necessity data you need to de-risk your assets and allocate resources efficiently. Contact our experts to initiate your target de-risking consultation today.
Creative Biolabs is your trusted partner in functional oncology discovery. By leveraging the precision of CRISPR-Cas9 screening and our deep expertise in complex cancer biology—from m6A epigenetics to the TME—we provide the high-confidence genetic dependency data required to accelerate your therapeutic pipeline and ensure your assets are focused on the most validated targets.
Contact Our Team for More Information and to Discuss Your Project
Reference
- Tan, Kezhe, et al. "CRISPR-Cas9 knockout screening identifies KIAA1429 as an essential gene in Ewing sarcoma." Journal of Experimental & Clinical Cancer Research 42.1 (2023): 250. https://doi.org/10.1186/s13046-023-02828-5. Distributed under Open Access license CC BY 4.0, without modification.
