Creative Biolabs offers customers a rapid, high quality TCR Vβ repertoire analysis service with a competitive price.
TCR Vβ repertoire analysis, which focus on analysis of TCRB genes or TCRB gene products (RNA, proteins, or both), is the most frequently used approach for TCR repertoire analysis, which plays an important role in studying the evolution process of TCR diversity during immune responses and the diagnosis of autoimmune diseases and immune deficiency.
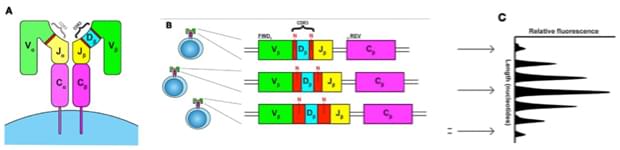 Fig.1 Unraveling the repertoire in wiskott-aldrich syndrome.1
Fig.1 Unraveling the repertoire in wiskott-aldrich syndrome.1
Characterizing TCR repertoires and acquiring knowledge on TCR amino acid sequence enable tracking of antigen-specific T cell clones, facilitate monitor T cell immunity and contributes to diagnosis and targeted therapy of T cell-related disorders. TCR sequencing involves PCR amplification and sequencing of the CDR3 region. And up to now, most TCR-sequencing has focused on TCRβ chain foe several reasons. Firstly, TCR β chain locus contains a D gene component while α chain locus is lack of it, TCR α chain locus only consists of V and J gene segments. So β chain has potential for greater combinatorial and junctional diversity than α chain. Secondly, each T cell expresses only a single β chain variant due to allelic exclusion mechanisms, which means that the number of β chain sequences is an indication of the number clonotypes. Finally, the TCR β chain tends to be the most heavily interrogated.
Our company has developed two different approaches for Vβ repertoire analysis: PCR analysis of TCRB genes and flow cytometric analysis of TCR Vβ families. The former one is based on detecting the presence or absence of the rearranged TCRB genes for studying the genotypes of TCR Vβ repertoire. The latter one adopts a panel of carefully selected Vβ specific monoclonal antibodies which can cover more than 70% of the whole TCR Vβ phenotypes.
Creative Biolabs is one of the few companies which offer the TCR Vβ repertoire analysis service. Compared with others, we can provide two analysis approaches and most professional and highest quality service just in order to meet all your research requirements.
To discuss your demands or to request a proposal, please contact us by .
Reference
For any technical issues or product/service related questions, please leave your information below. Our team will contact you soon.
All products and services are For Research Use Only and CANNOT be used in the treatment or diagnosis of disease.
 NEWSLETTER
NEWSLETTER
The latest newsletter to introduce the latest breaking information, our site updates, field and other scientific news, important events, and insights from industry leaders
LEARN MORE NEWSLETTER NEW SOLUTION
NEW SOLUTION
CellRapeutics™ In Vivo Cell Engineering: One-stop in vivo T/B/NK cell and macrophage engineering services covering vectors construction to function verification.
LEARN MORE SOLUTION NOVEL TECHNOLOGY
NOVEL TECHNOLOGY
Silence™ CAR-T Cell: A novel platform to enhance CAR-T cell immunotherapy by combining RNAi technology to suppress genes that may impede CAR functionality.
LEARN MORE NOVEL TECHNOLOGY NEW SOLUTION
NEW SOLUTION
Canine CAR-T Therapy Development: From early target discovery, CAR design and construction, cell culture, and transfection, to in vitro and in vivo function validation.
LEARN MORE SOLUTION