From Formulation Optimization to Targeting:
Aligning Liposome Parameters with Biological Readouts
Liposome formulation optimization is most effective when physicochemical parameters are evaluated alongside cellular uptake, biodistribution, payload release, and downstream biological performance. This page outlines how to connect formulation variables with the readouts that matter for candidate selection.
Why Liposome Formulation Optimization Often Fails to Predict Targeting
In lipid-based drug delivery, generating high-throughput data is no longer the primary hurdle; the bottleneck is identifying which formulation variables actually drive success. Formulation teams frequently optimize candidates strictly for uniformity—focusing on size, polydispersity index (PDI), and zeta potential. However, when these visually "perfect" liposomes advance to in vitro cell assays or in vivo models, they frequently fail to demonstrate intended targeting efficiency or functional outcomes.
Effective liposome formulation optimization starts by defining the biological endpoint first, then selecting formulation variables that are most likely to influence uptake, biodistribution, release behavior, and target engagement. To achieve this, formulation scientists must shift away from merely describing nanoparticles to predicting how their structure will withstand and utilize the biological milieu.
Key Takeaways for Successful Optimization
- • Size and zeta potential alone rarely explain targeting performance in complex physiological environments.
- • Lipid composition, PEG density, and internal buffer gradients primarily shape in vivo biological outcomes.
- • Formulation variables must be systematically screened against defined biological readouts, rather than optimized in isolation.
- • Design of Experiments (DoE) workflows help identify which structural parameters truly drive functional candidate performance.
What formulation optimization should really answer:
A complete screening logic connects physical changes to biological responses. It moves the project beyond "what is the liposome" to "what does the liposome do":
- Which variables control stability and protein corona formation?
- Which variables dictate specific cellular uptake?
- Which variables influence organ biodistribution?
- Which variables determine payload release and functional activity?
Which Liposome Parameters Commonly Drive Divergent Biological Readouts
Formulations with identical nanoscale profiles can yield radically different biological outputs. To build a highly functional delivery vehicle, developers must systematically evaluate a broader spectrum of parameters that dictate how a liposome interacts with complex physiological fluids. Here are the parameters that most commonly determine the ultimate fate of the formulation:
-
1. Lipid Composition & Molar Ratios
Altering the ratio of primary phospholipids to helper lipids (e.g., cholesterol) profoundly changes membrane rigidity and phase transition characteristics. This directly influences how serum proteins adsorb onto the surface, forming a unique protein corona that dictates subsequent cellular internalization pathways.
-
2. Surface PEG Density
While higher PEGylation improves in vivo circulation by shielding the liposome from immune recognition, excessive PEG density can sterically hinder active targeting ligands. Finding the precise "stealth vs. active" balance is a major driver of biodistribution.
-
3. Internal Buffer & Payload State
The internal pH gradient and buffer composition affect both encapsulation efficiency and the active pharmaceutical ingredient's (API) structural state. A robust internal environment acts as the primary driver for a predictable and triggerable release profile.
-
4. Ligand Density & Spacer Geometry
For targeted applications, it isn't just about having a ligand; it's about presentation. The ligand density and the length of the PEG spacer arm dictate whether the ligand can successfully protrude beyond the hydration layer to achieve multi-valent receptor binding, or if it will be obscured by surrounding lipids.
-
5. Particle Size Distribution & Serum Stability
A perfectly monodisperse sample in buffer might rapidly aggregate when exposed to serum. It is the size distribution under physiological conditions that truly governs whether the liposome will clear through the kidneys, be taken up by the liver, or successfully exploit leaky tumor vasculature.
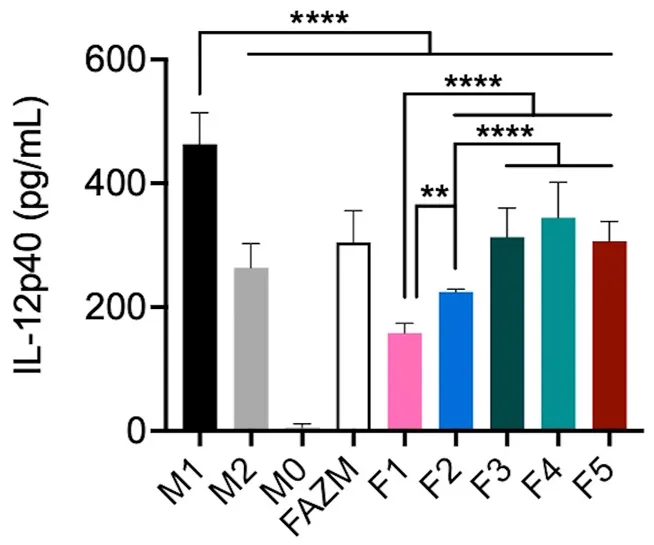
An Illustrative Example: Macrophage Activation
To demonstrate how precise parameter tweaking translates into biological function, consider research on Liposomal Azithromycin (L-AZM) formulations. While maintaining a consistent size and charge footprint, systematically modulating the lipid composition yielded drastically different macrophage responses. As seen in the study, optimized variations of the lipid molar ratio alone translated directly into specific, quantifiable levels of Interleukin-12 (IL-12) secretion, illustrating that biological alignment must supersede basic physical matching.
The "What to Measure" Checklist
The goal of characterization is not to generate more descriptors, but to identify which formulation attributes explain differences in uptake, targeting, biodistribution, and biological activity. A comprehensive optimization strategy requires analyzing metrics across both the physical structure and its biological interface. Here is a breakdown of what to measure and why each metric matters.
Formulation-Side Metrics
-
Particle Size & PDI:Determines the baseline clearance rate by the reticuloendothelial system (RES) and dictates the physical ability to extravasate into target tissues (e.g., via EPR effect).
-
Zeta Potential & Surface Charge:Influences colloidal stability and controls the degree of non-specific binding to negatively charged cell membranes or circulating proteins.
-
Encapsulation Efficiency (EE%):Directly dictates the therapeutic payload capacity and the economic viability of scaling the formulation for clinical translation.
-
Ensures the active pharmaceutical ingredient is held securely during circulation but releases efficiently upon reaching the specific target microenvironment.
-
Ligand Density & Orientation:Determines the receptor binding avidity; too few ligands lead to poor targeting, while too many can trigger immediate immune opsonization.
Biology-Side Readouts
-
Specific Cellular Uptake:Validates that the liposome is entering the correct cell type via the intended receptor-mediated pathway rather than random pinocytosis.
-
Endosomal Escape & Trafficking:Crucial for nucleic acids and certain proteins; confirms the payload avoids lysosomal degradation and successfully enters the cytosol.
-
Serum Stability & Protein Corona:Predicts in vivo half-life by demonstrating how the formulation resists premature breakdown and payload leakage in the presence of nucleases/proteases.
-
Organ Biodistribution:Proves organ-level targeting efficiency by tracking accumulation in the desired tissue versus clearance organs like the liver and spleen.
-
Target vs. Off-Target Binding Ratio:Defines the therapeutic window and safety profile, ensuring maximal efficacy with minimal systemic toxicity.
To ensure your data aligns correctly, leverage comprehensive formulation analysis and characterization. Advanced techniques like Cryo-TEM for morphology and SAXS for bilayer thickness provide the necessary depth to explain anomalous ex vivo and in vivo results.
A Practical Workflow for Liposome Formulation Optimization
Advancing a liposomal candidate from the bench to preclinical models requires a systematic framework. Rather than tweaking parameters via trial and error, a structured workflow connecting variables to defined biological endpoints accelerates candidate selection. We also emphasize that active targeting is not just a ligand question; it is a formulation question. Targeting strategies must be co-optimized with the lipid backbone and PEG shielding to succeed.
Define Endpoint
Determine the desired biological endpoint first (e.g., tumor accumulation, specific macrophage uptake, endosomal escape, or cytokine modulation).
Select Variables
Select the formulation variables most likely to influence that specific endpoint (lipid molar ratio, cholesterol content, PEG density, ligand density, internal buffer, or release profile).
Characterize
Use orthogonal characterization to confirm structural differences across candidates (DLS + zeta + morphology + encapsulation efficiency + release kinetics + serum stability).
Map & Rank
Map each formulation to biological readouts and rank the candidates based on real performance (cellular uptake, receptor engagement, biodistribution, target/off-target ratio, functional response).
Why this workflow accelerates translational selection: Relying solely on One-Factor-At-A-Time (OFAT) screening often masks synergistic effects—for example, how high cholesterol content might offset the stabilizing benefits of a longer PEG chain. Implementing Design of Experiments (DoE) workflows combined with orthogonal characterization establishes a robust predictive model. This approach minimizes guesswork, ensuring that when a candidate enters expensive in vivo testing, its physicochemical properties are already mathematically linked to the highest probability of biological success.
Parameter-to-Biology Matrix
| Formulation Parameter | Physicochemical Effect | Aligned Biological Readout |
|---|---|---|
| Lipid Composition & Molar Ratio | Alters membrane fluidity, structural rigidity, and influences protein corona adsorption patterns. | In vivo organ biodistribution, specific macrophage/cellular activation pathways. |
| Surface PEGylation Density | Increases steric hindrance and hydration layer thickness, preventing opsonization. | In vivo circulation half-life, reduction in non-specific in vitro phagocytosis. |
| Internal Buffer & pH Gradient | Dictates active loading efficiency and the intravesicular precipitation state of the drug. | Ex vivo therapeutic window, mitigation of premature systemic toxicity. |
| Targeting Ligand Conjugation | Modifies surface chemistry and introduces accessible specific receptor binding sites. | Target-to-off-target accumulation ratio, enhanced in vitro receptor-mediated endocytosis. |
Optimize Formulation Variables
Let our experts guide your candidate selection with data-driven matrix modeling and DoE strategies.
Request Optimization ServiceDevelop Targeted Systems
Co-optimize targeting ligands, spacer lengths, and the lipid backbone to ensure precise in vivo delivery.
Targeted Liposome ServicesFrequently Asked Questions
While particle size and zeta potential are important baselines, the most critical parameters for predicting biological performance include lipid composition (which dictates membrane fluidity and protein corona formation), surface PEG density, targeting ligand geometry, and the internal buffer environment that controls drug release kinetics.
Evaluating targeting efficiency requires moving beyond static characterization. It involves pairing the formulation data with dynamic biological assays, measuring specific cellular uptake against non-specific phagocytosis, assessing target-to-off-target accumulation ratios in in vivo biodistribution studies, and verifying downstream functional responses like specific cytokine modulation.
Formulations often fail in vivo because they were optimized strictly for uniformity or stability in buffer solutions, rather than functional efficacy in complex physiological fluids. Issues like premature burst release in circulation, immediate opsonization by immune proteins, or steric hindrance of targeting ligands by excessive PEG chains can render a structurally "perfect" liposome biologically ineffective.
Design of Experiments (DoE) replaces the inefficient "one-variable-at-a-time" approach by allowing researchers to simultaneously test multiple parameters (e.g., lipid ratios, internal pH, ligand density) and analyze their interactions. This statistical modeling identifies which specific variables—and variable combinations—truly drive the desired biological readout, accelerating candidate selection.
Orthogonal characterization should be paired with biological assays such as flow cytometry for cellular internalization, confocal microscopy for endosomal trafficking, serum stability testing to map protein corona formation, and targeted functional assays (like gene silencing or protein expression mapping) to verify the payload is successfully delivered and active.
References
- Masud, Abdullah A., et al. "Optimization and Characterization of a Liposomal Azithromycin Formulation for Alternative Macrophage Activation." Frontiers in drug delivery 2 (2022): 908709. https://doi.org/10.3389/fddev.2022.908709
- Under Open Access license CC BY 4.0, without modification. CC BY 4.0
Online Inquiry
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.