Optimizing Ligand Density on Liposomes
for Targeted Delivery
Learn how ligand density influences receptor binding, PEG shielding, non-specific uptake, and internalization behavior in targeted liposome design.
The Core Challenge of Ligand Density on Liposomes
In the realm of precision nanomedicine, active targeting liposomes represent a significant leap forward from traditional passive delivery methods relying solely on the Enhanced Permeability and Retention (EPR) effect. However, a pervasive and highly complex challenge plagues formulation scientists: determining the optimal surface density of targeting ligands.
Researchers often successfully identify a highly specific disease-relevant receptor (such as HER2 in breast cancer or Folate receptors in various malignancies) and carefully select an appropriate high-affinity ligand. Yet, the translation of this pairing into an effective liposomal formulation frequently falters at the engineering stage. In targeted liposome design, increasing ligand loading does not necessarily improve delivery performance. Beyond a certain threshold, higher ligand density may reduce accessibility, disrupt stealth properties, and increase off-target interactions.
Developing an effective platform starts with precise Targeting Ligand Selection and Design, but the critical next step is establishing the exact density. Setting the ligand density too low results in insufficient binding avidity, leading to poor target cell recognition and minimal endocytosis. Conversely, there is no universal optimal ligand density; the working range must be defined experimentally for each ligand–receptor–formulation system.
How Much Ligand Should Be Added to a Targeted Liposome?
Understanding the delicate interplay between receptor affinity and steric hindrance is paramount. When multiple ligands on a single liposome simultaneously engage multiple receptors on a cell surface, they create a highly stable multivalent interaction known as "avidity." This thermodynamic "velcro effect" is the primary goal of liposome surface functionalization.
However, actual surface accessible ligand density is rarely equivalent to the theoretical feed ratio. Biological receptors exist within a complex, crowded glycocalyx. If ligands are situated too densely on the liposome, they compete with one another for space, leading to steric hindrance. Ligand accessibility is heavily influenced by PEG spacer length, conjugation efficiency, ligand orientation, and surface segregation. Furthermore, excessive functionalization can collapse or compress the essential polyethylene glycol (PEG) corona, which shields PEGylated liposomes from rapid systemic clearance.
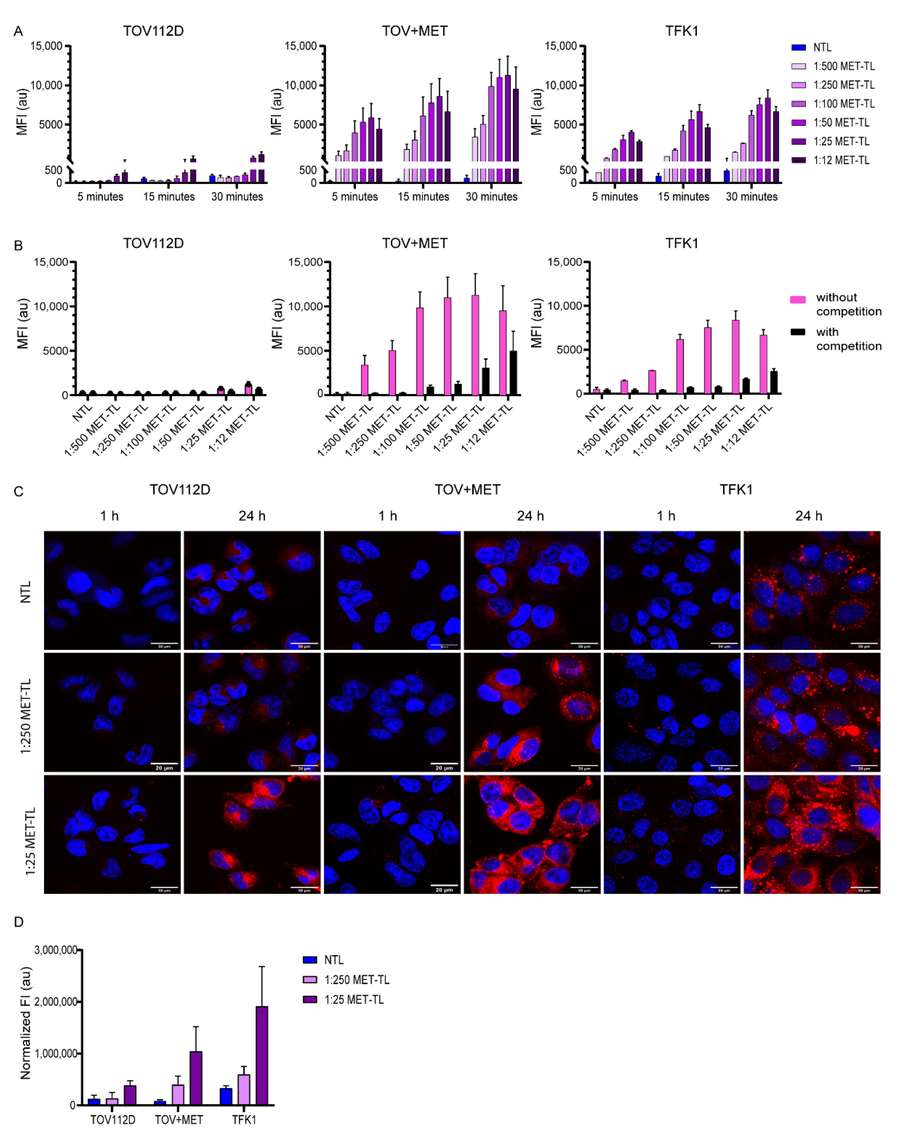
Scientific Context: As highlighted in recent literature, the image illustrates a typical targeted-liposome logic: selecting a disease-relevant receptor, choosing a matching ligand, conjugating that ligand onto the liposome surface, and then confirming whether the modification translates into stronger cellular uptake. Finding the exact density is what turns theory into therapeutic efficacy.
Why Can Excessive Ligand Density Alter Internalization Pathways?
The density of surface ligands not only influences whether a liposome will bind to a cell but can also affect receptor-mediated endocytosis. Cellular uptake mechanisms—predominantly clathrin-mediated endocytosis, caveolae-mediated endocytosis, and macropinocytosis—are sensitive to the physical properties of the binding event.
Higher ligand density may enhance receptor clustering and alter membrane remodeling during uptake. Depending on receptor biology and cell context, this can change internalization behavior, but the dominant pathway should be verified experimentally rather than assumed from density alone. A liposome with a low density of ligands might achieve sufficient avidity to induce membrane invagination via clathrin-dependent pathways, whereas extensive cross-linking of cell-surface receptors by densely packed ligands can bias the system towards alternative routes.
Crucially, the chosen endocytic pathway determines the intracellular fate of the therapeutic payload. Clathrin-mediated pathways quickly route to acidic endolysosomal compartments—ideal for pH-sensitive liposomes but potentially destructive to sensitive biologics unless robust endosomal escape mechanisms are incorporated. Understanding this relationship ensures that in vitro binding assays accurately predict in vivo functional delivery.
A Practical Workflow for Ligand Density Optimization
Formulation scientists should approach targeted nanoparticle uptake by treating ligand density as an empirical variable. A structured, data-driven workflow is essential for finding the optimal formulation.
Fix Core Parameters
Standardize liposome size, total PEG content, and ligand chemistry before varying conjugation density.
Prepare Density Series
Formulate a screening panel of varying ligand densities rather than a single theoretical construct.
Quantify Incorporation
Measure the true surface-accessible ligand density, discarding theoretical feed ratios.
Evaluate Specific Binding
Compare binding affinity on receptor-high versus receptor-low (or negative) cell lines.
Measure Internalization
Assess actual intracellular delivery and functional payload release, not just surface binding.
Test Serum Stability
Expose liposomes to serum-containing media to evaluate protein corona sensitivity and stealth property retention.
Assess Phagocytic Liability
Evaluate macrophage uptake in vitro to predict clearance rates by the mononuclear phagocyte system.
Select Optimal Density
Base the final decision on total delivery performance, therapeutic index, and safety profile, not merely on maximum receptor binding.
How to Determine the Optimal Ligand Conjugation Density?
Achieving targeted delivery requires rigorous, iterative Liposome Formulation Optimization to screen the molar ratios of functionalized lipids against background matrix lipids. Start with a screening panel rather than a fixed optimum.
| Density Level | Target Binding (Avidity) | Non-Specific Uptake | Circulation Half-life | Cellular Internalization |
|---|---|---|---|---|
| Low (<0.5 mol%) | Weak (Insufficient) | Very Low | Long (Intact PEG) | Poor |
| Common screening range* | Strong (Optimal) | Low | Prolonged | High (Receptor-mediated) |
| High (>5 mol%) | Reduced (Steric Hindrance) | High (Opsonization) | Short (Rapid Clearance) | Variable (Often Non-specific) |
How Should Ligand Density Be Validated Experimentally?
Theoretical feed ratios should not be treated as equivalent to surface-accessible ligand density. Experimental quantification is needed to confirm conjugation efficiency, ligand accessibility, and functional performance. Comprehensive Formulation Analysis and Characterization ensures that the targeted liposome operates as intended both ex vivo and in vivo.
A. Quantifying Actual Surface Density
- • Accessible Ligand Quantification: Use fluorescence labeling, HPLC, or amino acid analysis to determine true post-insertion conjugation efficiency.
- • Binding Kinetics: Utilize Surface Plasmon Resonance (SPR) or ELISA-like binding readouts to assess the structural orientation and avidity of the entire liposomal construct.
- • Physicochemical Assessment: Track changes in Dynamic Light Scattering (DLS) and Zeta potential to ensure high ligand density hasn't caused particle aggregation or altered surface charge significantly.
B. Proving Receptor-Mediated Uptake
- • Receptor Specificity Controls: Perform flow cytometry and confocal microscopy on both receptor-positive and receptor-low/negative cell lines to verify specific targeting.
- • Competition Assays: Introduce free targeting ligands to block receptors prior to liposome administration to confirm that cellular internalization is genuinely receptor-mediated.
- • Serum-Condition Testing: Evaluate uptake in serum-containing media to ensure the ligand-receptor axis is not completely obstructed by the in vivo protein corona.
Frequently Asked Questions
PEG content and ligand density are highly interdependent. It is generally recommended to fix the baseline stealth properties (e.g., 2-5 mol% PEG) first to establish basic in vivo stability, and then titrate the ligand-conjugated lipid. Alternatively, using a Design of Experiments (DoE) approach allows scientists to co-optimize both variables simultaneously to map the complete interactive design space.
Directly comparing small peptides and full monoclonal antibodies by lipid molar percentage is misleading due to drastic differences in molecular weight and footprint. A 1 mol% conjugation of a bulky antibody will cause severe steric crowding, whereas 1 mol% of a small peptide may leave ample room on the liposome surface. Density should often be evaluated as "ligands per liposome particle" to make meaningful cross-ligand comparisons.
Maximum receptor binding (avidity) does not guarantee successful internalization. Extremely high affinity, caused by dense ligand clustering, can cross-link receptors to the point where membrane invagination and endocytosis are physically stalled. Furthermore, if a liposome is "stuck" to the cell surface too strongly, it may not proceed to endosomal sorting or payload release.
PEG (polyethylene glycol) is applied to liposomes to create a steric hydration layer that prevents plasma proteins from binding. When ligand density is too high, the bulky ligands can physically disrupt this PEG corona. This exposes the hydrophobic lipid bilayer underneath, causing rapid opsonization (protein binding) and subsequent clearance by the immune system, effectively negating the stealth benefits.
Yes, significantly. If the ligand is conjugated directly to the lipid surface without a spacer, it will be buried underneath the PEG corona and remain invisible to the target cell receptor. Ligands are typically conjugated to the distal end of a PEG spacer (e.g., PEG2000 or PEG3400) to ensure they extend beyond the protective hydration layer and can freely interact with the biological environment.
References
- Mesquita, Bárbara S., et al. "The impact of nanobody density on the targeting efficiency of PEGylated liposomes." International Journal of Molecular Sciences 23.23 (2022): 14974. https://doi.org/10.3390/ijms232314974
- Under Open Access license CC BY 4.0, without modification.
Online Inquiry
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.