Imaging Liposome Development: Balancing Labeling Strategy, Signal-to-Noise Ratio, and Stability
Imaging liposome development requires more than adding a detectable probe. Effective formulations must support clear imaging signals while preserving liposome integrity, minimizing background noise, and maintaining biologically relevant behavior in preclinical studies.
Key Considerations in Imaging Liposome Development
Successful imaging liposome development begins with choosing a labeling approach that aligns with your downstream pharmacological goals. For biopharma researchers and formulation scientists, the visualization of lipid-based nanoparticles is crucial for interpreting biodistribution analysis and executing real-time in vivo tracking. However, creating these functionalized vesicles presents a profound, multidimensional challenge.
The central difficulty is that introducing a detectable probe into a meticulously balanced lipid formulation often causes membrane perturbation, altered circulation profiles, label dissociation, and potentially false biodistribution readouts. A major goal in imaging liposome development is to preserve liposome behavior after probe integration.
A highly emissive probe that improves visibility may compromise membrane integrity, increasing the risk of premature payload leakage. Conversely, an overly conservative formulation might remain physically stable but fail to generate the robust, interpretable data required for critical in vivo evaluations. Researchers must carefully navigate these variables—labeling strategy, signal-to-noise ratio (SNR), and formulation stability—to yield biologically meaningful pharmacokinetic data without disrupting the intrinsic behavior of the nanocarrier.
The Triple Balance Paradigm
-
1
Labeling Strategy: Selecting the correct modality and placement based on experimental demands.
-
2
Signal-to-Noise Ratio: Minimizing background autofluorescence and clearance-induced artifacts to ensure accurate representation.
-
3
Formulation Stability: Preventing probe-induced lipid phase separation, aggregation, or premature release during circulation.
Recommended Decision Framework for Imaging Liposome Development
Navigating the balance between signal and stability often requires a practical decision matrix. The framework below aligns research goals with appropriate labeling strategies and their necessary mitigations.
| Research Goal | Preferred Labeling Strategy | Main SNR Risk | Main Stability Risk | Practical Mitigation |
|---|---|---|---|---|
| Short-term in vitro uptake | Bilayer fluorescent dye | Self-quenching / nonspecific signal | Membrane perturbation | Keep dye to a low mol%; meticulously verify size and PDI. |
| Whole-body biodistribution | PET/SPECT surface chelation or remote loading | Free isotope background from transchelation | Altered surface behavior & MPS clearance | Extensive serum stability testing and rigorous radiochemical purity checks. |
| Deep-tissue longitudinal tracking | Radionuclide / MRI contrast agents | Low sensitivity for some MRI agents | Payload leakage / nanoparticle aggregation | Orthogonal validation comparing functional imaging with physicochemical stability. |
Labeling Strategies for Imaging Liposome Development
Instead of merely introducing imaging modalities, researchers must choose them based on three distinct dimensions: required imaging depth and duration, whether intact liposomes or released cargo must be distinguished, and the formulation's tolerance for surface modification.
Fluorescent probes remain highly effective for in vitro and ex vivo analyses due to their high resolution and multiplexing capabilities. Near-Infrared (NIR) dyes (e.g., Cy5.5, ICG) are typically preferred to bridge the gap to in vivo tracking, offering improved tissue penetration and reduced optical scattering. For customized fluorescent optimization, leveraging a specialized Fluorescent Liposome Development Service can streamline preclinical screening.
For translational and clinical studies, nuclear imaging (PET/SPECT) and MRI are often employed. Radionuclides (e.g., 99mTc, 111In) can be attached via surface chelators or utilizing "after-loading" techniques.
Crucially, the placement of the label dictates both performance and stability:
- Aqueous core loading: Generally preferred for minimizing surface interactions, though it often requires active loading mechanisms.
- Lipid bilayer insertion: Straightforward for lipophilic dyes, but can increase the risk of leakage by disrupting lipid packing density.
- Surface conjugation: Highly effective for radiometals, yet it may compromise the stealth properties of the PEG layer, inviting premature immune clearance.
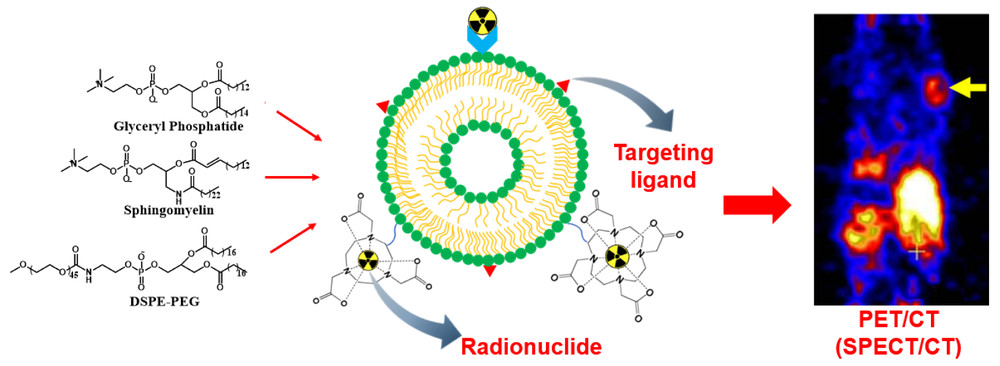
Insight from Literature
This scheme illustrates that imaging liposome development is not only about adding a detectable label. The imaging moiety, targeting ligand, and surface engineering must be integrated in a way that preserves circulation behavior and formulation integrity. In practice, stronger detectability does not automatically mean better performance, because excessive or poorly positioned labeling may alter biodistribution, increase background signal, or reduce liposome stability.
Balancing Signal and Stability in Imaging Liposome Development
A critical hurdle is interpreting the signal accurately against a complex biological background. A strong signal is only valuable if it genuinely reflects the location of the intact liposome.
Sources of Background Signal
Background artifacts primarily arise from probe leakage, free dye clearance through renal pathways, tissue autofluorescence, and nonspecific macrophage uptake.
When probes leak into systemic circulation, they often create a "false positive" background that misrepresents the actual nanocarrier distribution.
How to Improve Interpretability
To enhance SNR, formulators should choose appropriate NIR wavelengths, rigorously remove unincorporated probes via Size Exclusion Chromatography (SEC) or dialysis, and test retention in serum.
Including unlabeled and free probe controls in vivo is highly recommended to establish true baseline comparisons.
Why Strong Signal Can Mislead
Do not equate "seeing a signal" with "seeing an intact liposome." Released probes distribute differently than intact vesicles.
Furthermore, phenomena like fluorophore quenching/dequenching, or surface-bound labels altering the intrinsic physiological fate, can produce highly misleading data.
To navigate these complexities and mathematically optimize your formulation, expert guidance is often required.
Explore Imaging Liposome Development ServicesSafeguarding Formulation Stability
The introduction of imaging moieties acts as a structural impurity within the highly organized lamellar phase of the liposome. In imaging liposome development, stability is multi-dimensional:
- Physical stability: Ensuring minimal changes in vesicle size, preventing aggregation, and maintaining a low polydispersity index (PDI).
- Chemical stability: Protecting against lipid oxidation and maintaining the chemical integrity of the imaging probe.
- Label retention stability: Ensuring the imaging agent remains tightly associated with the liposome under physiological shear and protein interactions.
Bulky fluorophores embedded in the bilayer can alter the gel-to-liquid crystalline phase transition temperature (Tm). This disruption frequently leads to increased membrane fluidity and accelerated payload leakage. To counter this, adjusting the molar ratios of high-Tm lipids and cholesterol is typically preferred to re-rigidify the membrane. Continuous assessment through a rigorous Formulation Stability Monitoring Service provides essential verification over extended periods.
The Role of Cholesterol
Introducing lipophilic dyes often disrupts the tight packing of phospholipids. Cholesterol acts as a critical fluidity buffer, filling the interstitial spaces between lipid tails. By carefully titrating the cholesterol ratio, formulators can re-stabilize the membrane and prevent the early leakage of both the imaging probe and encapsulated therapeutic payloads, a step that is central to successful imaging liposome development.
Translational Characterization Protocols
Validating an imaging liposome requires a multi-tiered approach. Formulation scientists must confirm that the labeled liposome behaves identically to its unlabeled parent counterpart.
| Parameter Evaluated | Primary Objective in Imaging Context |
|---|---|
| Post-labeling Size/PDI Comparison | Compare directly with the unlabeled parent liposome to ensure labels do not induce swelling or aggregation. |
| Free Probe Removal Confirmation | Verify that unbound imaging agents have been completely separated prior to in vivo injection to avoid false backgrounds. |
| Probe Retention after Serum Incubation | Test leakage in serum-containing media to confirm probe retention under simulated in vivo physiological conditions. |
| Signal Linearity vs. Concentration | Ensure that the imaging signal remains proportional to liposome concentration, avoiding artifacts like fluorophore self-quenching. |
| Intact Liposome vs. Released Probe | Implement assays (e.g., FRET or dual-isotope tracking) to actively discriminate the intact nanocarrier from free molecules. |
Orthogonal confirmation using both imaging readouts and rigorous physicochemical assays is the safest route to clinical translation.
Access Basic Characterization ServicesConclusion: Optimizing Imaging Liposome Development
For translational imaging liposome development, success is most readily achieved when the labeling strategy, signal quality, and formulation stability are optimized together rather than independently. The most informative formulations are not necessarily those with the strongest raw signal, but those that successfully preserve native liposome behavior while generating interpretable, robust imaging readouts in complex physiological environments.
Frequently Asked Questions
In imaging liposome development, the acceptable labeling threshold depends heavily on the probe's lipophilicity and the bilayer composition. For lipid-anchored fluorophores, exceeding 1-2 mol% often induces significant membrane perturbation, leading to fluorophore self-quenching and payload leakage. Iterative probe-to-lipid ratio optimization is necessary to identify the maximum signal threshold that does not alter physical stability.
This depends on your specific imaging modality and the liposome's payload. Aqueous core encapsulation (e.g., loading water-soluble dyes or specific radionuclides) is often preferred to preserve the exterior lipid membrane integrity and circulation half-life. However, bilayer insertion is straightforward for lipophilic probes and provides high labeling efficiency, though it requires careful monitoring of lipid packing disruptions.
It is crucial not to equate raw signal with intact liposomes. To verify, researchers employ techniques like Förster Resonance Energy Transfer (FRET) or dual-isotope tracking where a signal shift occurs upon vesicle rupture. Additionally, parallel ex vivo tissue homogenization paired with chromatographic analysis (SEC) can definitively separate intact nanocarriers from free, released dye molecules.
Rigorous controls include the unlabeled parent liposome (to verify that physical parameters like size and PDI remain unchanged) and a "free probe" control injected in vivo (to map the baseline biodistribution of the unencapsulated imaging agent). These controls are indispensable for isolating true liposomal tissue accumulation from background noise and physiological clearance pathways.
For high-throughput, short-term screening (e.g., in vitro cell uptake or superficial ex vivo analysis), fluorescence imaging is highly effective due to its resolution and ease of multiplexing. For translational whole-body biodistribution, nuclear imaging (PET/SPECT) is the preferred choice as it provides fully quantitative, deep-tissue, 3D tomographic tracking without the limitations of optical scattering.
References
- Low, Ho Ying, et al. "Radiolabeled liposomes for nuclear imaging probes." Molecules 28.9 (2023): 3798. https://doi.org/10.3390/molecules28093798
- Under Open Access license CC BY 4.0, without modification.
Online Inquiry
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.