Why High-Affinity Aptamers Do Not Always Translate into Efficient Liposomal Delivery
A systematic framework for identifying bottlenecks in targeted liposome delivery, from ligand orientation and PEG shielding to cellular uptake and intracellular payload release.
Efficient Delivery is a Biological Cascade
High affinity improves the probability of target engagement, but delivery success depends on a multistep biological cascade from binding to intracellular release.
Delivery scientists frequently identify a candidate aptamer demonstrating nanomolar or even picomolar binding affinities to a target receptor in vitro. However, high binding affinity measured for a free aptamer often fails to translate into efficient liposomal delivery after surface conjugation. The outcome is characterized by excellent surface binding but poor cellular internalization, suboptimal payload release, and ultimately, an efficacy that falls short in vivo.
This disconnect requires researchers to diagnose the bottleneck systematically: is the problem inherent to the aptamer itself, the lipid formulation, the spatial orientation of the ligand, the PEG shielding dynamics, or the intracellular transport pathways? Translating molecular affinity into a targeted nanomedicine requires separating target recognition from productive cellular uptake.
Furthermore, the nature of the payload heavily dictates the stringency of these requirements. Small-molecule delivery may tolerate limited endosomal escape, as many chemotherapeutics can passively diffuse across lipid membranes once the liposome degrades in the lysosome. Conversely, nucleic acid payloads (like siRNA or mRNA) are much more dependent on productive cytosolic release; failure to escape the endosome results in complete therapeutic failure.
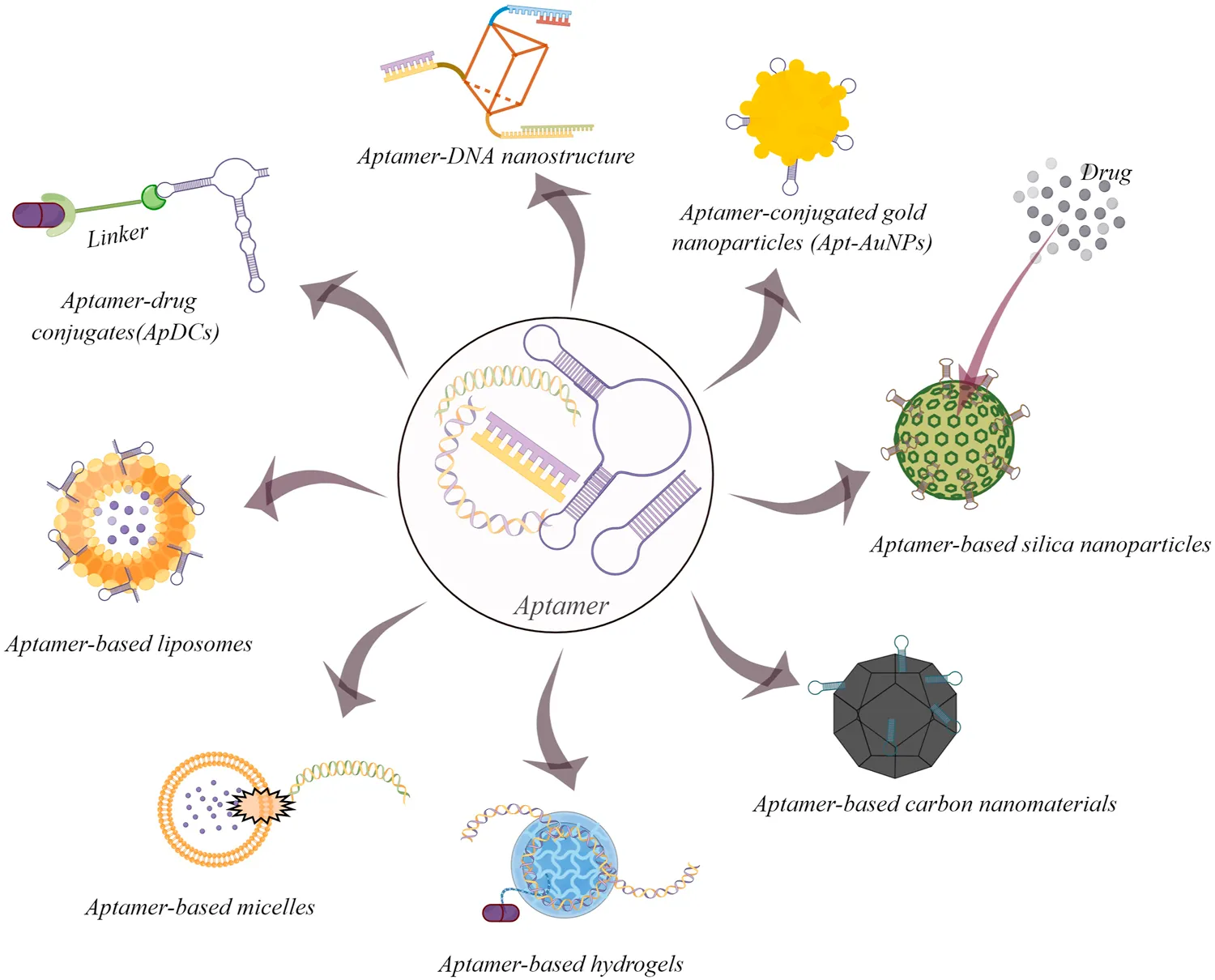
Fig.1 Some common examples of aptamer-based drug delivery systems for cancer therapy.1,2
The picture highlights the versatility of aptamers as targeting elements across multiple drug delivery platforms. For aptamer-modified liposomes, the key takeaway is not simply that aptamers bind well, but that successful delivery still requires converting target recognition into productive cellular uptake and payload transport.
Assessing the Aptamer Itself: Structure, Biology, and Environment
Before altering lipid compositions, it is crucial to determine if the aptamer maintains its functionality when transitioning from a purified buffer environment to the surface of a liposome in a complex biological matrix.
Conformation and Target Accessibility
Aptamers selected against purified recombinant proteins frequently fail to recognize the native cell-surface conformation of the receptor. Furthermore, an epitope highly accessible in vitro may be sterically hindered by the glycocalyx or other membrane proteins on live cells.
Even if binding occurs, the targeted receptor may be a static structural protein that does not readily undergo endocytosis. Strong binding without subsequent receptor internalization effectively anchors the liposome to the plasma membrane, preventing payload delivery.
Attachment Site and Stability
Conjugation chemistry can inadvertently destroy aptamer function. Attaching a lipid anchor to the wrong terminus (5' vs 3') can disrupt the delicate secondary structures (e.g., G-quadruplexes or hairpins) essential for binding. Moreover, aptamer affinity measured in simple buffers often drops precipitously in serum due to nuclease degradation or competitive binding by serum proteins.
To mitigate these risks and ensure the aptamer remains active post-conjugation, researchers rely on specialized Aptamer-Modified Liposome Development services to evaluate the optimal linker strategy and secondary structure preservation.
Systematic Formulation and Engineering Matrix
The physical and chemical composition of the liposome directly modulates the performance of the targeting ligand. Precise Liposome Formulation Optimization is required to balance pharmacokinetics with active targeting capabilities.
| Parameter | Why It Matters | Common Failure Mode | Optimization Consideration |
|---|---|---|---|
| Receptor Biology | Determines whether a binding event triggers active endocytosis. | Strong surface binding but weak or stalled cellular uptake. | Prioritize targets known to mediate clathrin/caveolae-dependent internalization. |
| Aptamer Attachment Site | Controls spatial orientation and preserves specific folding. | Loss of target affinity immediately after lipid conjugation. | Systematically screen 5' versus 3' modifications and evaluate post-conjugation SPR. |
| Spacer Length | Maintains physical distance between the lipid bilayer and the aptamer binding pocket. | Buried ligand; steric hindrance prevents receptor interaction. | Compare varying carbon chain or PEG lengths for the aptamer anchor. |
| Ligand Density | Balances multivalent avidity against steric crowding and immune clearance. | Competitive self-inhibition on the liposome surface; rapid clearance in vivo. | Conduct DOE titration (typically 0.1% to 2 mol%) to locate the optimal uptake density. |
| PEG Architecture | Extends circulation half-life but risks masking the targeting ligand. | The "PEG Dilemma": long circulation but complete failure of active targeting. | Utilize differential PEG layers (e.g., PEG3400 for aptamer tether, PEG2000 for shielding). |
| Lipid Composition | Influences membrane fluidity, stability, and endosomal escape capability. | Successful uptake but payload remains trapped and degrades in lysosomes. | Introduce pH-sensitive ionizable lipids or fusogenic lipids (e.g., DOPE). |
| Serum Stability | Dictates integrity of the RNA/DNA sequence during systemic circulation. | Rapid loss of targeting function in vivo due to nuclease digestion. | Incorporate 2'-fluoro or 2'-O-methyl chemical modifications into the sequence. |
Recommended Evaluation Workflow for Aptamer-Modified Liposomes
Before applying a troubleshooting framework, researchers must establish a robust analytical pipeline to identify exactly where the delivery process stalls. We recommend a sequential five-step evaluation workflow to pinpoint the bottleneck:
Free Aptamer Binding
Do not rely solely on SPR. Use flow cytometry to confirm the fluorescent free aptamer binds robustly to native receptors on live target cells.
Binding Retention
Assess the liposome-aptamer conjugate via flow cytometry. A severe drop in affinity indicates conjugation-induced steric hindrance.
Cellular Uptake Quantification
Differentiate surface binding from endocytosis. Use quenching assays (e.g., trypan blue) to quantify successful internalization of the liposomes.
Endosomal Colocalization
Use confocal microscopy and LysoTracker to determine if nanocarriers are trapped in lysosomes or successfully achieve endosomal escape.
Functional Payload Readout
Evaluate therapeutic efficacy (e.g., gene silencing) in media containing 10-50% serum to bridge the gap toward physiological reality.
A Practical Troubleshooting Framework
Once you have gathered the data from the evaluation workflow, apply this stepwise analytical approach. This decision tree helps determine whether to reconsider the biological target, the ligand sequence, or the lipid formulation parameters.
Target binding is weak or absent.
If the free aptamer binds but the conjugated liposome does not, the conjugation process is likely the culprit. This indicates steric masking, loss of secondary structure, or improper spatial orientation. A shift in attachment strategy (e.g., moving from the 5' to the 3' end) or employing a longer PEG spacer is required.
Binding is confirmed, but cellular uptake is poor.
Do not assume that target binding equates to endocytosis. If liposomes remain on the cell surface, the targeted receptor may not actively mediate internalization. Alternatively, the ligand density may be suboptimal. Titrate the ligand density; too few aptamers yield weak avidity, while too many cause competitive inhibition.
Uptake is successful, but there is no therapeutic effect.
If confocal microscopy confirms internalization but payload efficacy is absent, the payload is likely trapped in endo-lysosomal compartments. This requires re-engineering the lipid matrix for intracellular routing, leveraging technologies like ATP-Responsive Liposome Development or utilizing fusogenic helper lipids to force cytosolic release.
Why Strong In Vitro Performance Still Fails In Vivo
A recurring challenge for preclinical project leads and formulation scientists is the sharp decline in efficacy when transitioning from cell culture to animal models. Several critical translation risks must be proactively addressed before advancing to in vivo studies:
Protein Corona Masking
Upon intravenous injection, liposomes are immediately coated by a complex layer of serum proteins. This protein corona can physically bury the aptamer, completely neutralizing its targeting capability before it reaches the target tissue.
Nuclease Degradation
Unmodified RNA or DNA aptamers that perform well in controlled in vitro assays are highly vulnerable to rapid enzymatic cleavage by endogenous nucleases in the bloodstream, leading to functional loss.
Altered Biodistribution and RES Uptake
Even perfectly targeted liposomes must evade the reticuloendothelial system (RES). Excessive aptamer density or suboptimal PEG shielding can inadvertently increase opsonization, leading to rapid clearance by the liver and spleen rather than accumulation at the tumor site.
Different Receptor Accessibility
The 3D architecture of solid tumors presents barriers absent in 2D cell cultures. Dense extracellular matrix (ECM), high interstitial fluid pressure, and heterogeneous receptor expression mean that an epitope highly accessible on cultured cells may be physically unreachable in actual tissue.
Ensure your formulation is ready for the next phase by evaluating it across relevant biological matrices.
Access Formulation Stability Monitoring ServicesFrequently Asked Questions
The primary suspect is usually steric hindrance or loss of aptamer secondary structure during the chemical conjugation process. Verify that the attachment site (5' vs 3') does not disrupt the folding geometry of the binding pocket. Additionally, check your PEG architecture: if the aptamer is conjugated to a short linker while the liposome is heavily shielded with long PEG chains, the ligand is likely buried and physically cannot reach the cellular receptor.
This distinction requires precise imaging and functional assays. Use dual-labeled liposomes (e.g., fluorescent lipid and a differently tagged payload) paired with confocal microscopy. If you observe strong fluorescence accumulated strictly at the cell membrane outline, internalization is failing. If fluorescence appears as punctate dots inside the cell but no biological readout (e.g., gene silencing) is achieved, the liposomes are successfully internalizing but failing to execute endosomal escape.
A standard optimization range typically falls between 0.1% and 2 mol% of total lipids. Starting around 0.5 mol% is a common baseline. Density must be titrated carefully: too sparse, and you fail to achieve the multivalent avidity needed to trigger endocytosis; too dense, and you risk steric crowding that causes ligands to physically block each other from engaging receptors, alongside increasing the risk of rapid immune clearance in vivo.
Always validate the aptamer's functionality against live, relevant cells first. Confirm that the sequence can bind native receptor conformations and trigger endocytosis when attached to a simple fluorescent reporter. Once the biology of the ligand is validated, lock the sequence and transition entirely to optimizing the liposome formulation (lipid ratios, spacer lengths, PEG density) to construct a successful delivery vehicle.
Upon intravenous injection, liposomes rapidly adsorb serum proteins, forming a "protein corona." This layer can physically mask the aptamers, completely negating their targeting capability. Additionally, unmodified nucleic acid aptamers are susceptible to rapid degradation by endogenous nucleases. Proper chemical modifications and careful PEG shielding strategies are essential to preserve the ligand's function during systemic circulation.
References
- Gao, Fei, et al. "Recent advances in aptamer-based targeted drug delivery systems for cancer therapy." Frontiers in bioengineering and biotechnology 10 (2022): 972933. https://doi.org/10.3389/fbioe.2022.972933
- Under Open Access license CC BY 4.0, without modification.
Online Inquiry
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.