Anti-Sulfotransferase Antibody Development Service
Sulfotransferases (SULTs) are a critical family of enzymes that catalyze the sulfation of various substrates, including hormones, neurotransmitters, and drugs, as well as complex carbohydrates such as glycosaminoglycans and glycolipids. In the context of protein engineering and structural biology, antibodies targeting these enzymes serve not only as detection tools but also as sophisticated model systems for studying protein-protein interactions. Creative Biolabs provides a comprehensive Anti-Glycan Related Enzyme Antibody Development Service designed to generate high-affinity binders against sulfotransferases. Our platform is particularly optimized for research teams utilizing anti-sulfotransferase single-chain variable fragments (scFv) to investigate strategies for generating affinity-matured mutants through targeted mutation of the VH framework region 1 (FR1).
The Role of Sulfotransferases and Antibodies in Protein Engineering
Sulfotransferases are broadly categorized into cytosolic SULTs, which metabolize small molecules, and membrane-associated Golgi sulfotransferases, which modify glycans and proteins. While their biological function in detoxification and signaling is well documented, their structural complexity makes them excellent candidates for antibody engineering studies. An anti-sulfotransferase antibody, particularly in the scFv format, provides a robust scaffold for exploring how changes in non-CDR regions influence binding kinetics.
Research has demonstrated that mutations in the framework regions (FRs), specifically the VH framework region 1, can significantly alter the conformational stability and flexibility of an antibody variable domain. This "remote control" mechanism allows for affinity maturation without directly altering the antigen-binding loops. By developing a diverse panel of antibodies against specific sulfotransferase targets—such as heparan sulfate sulfotransferases or tyrosylprotein sulfotransferases—researchers can create a library of mutants to dissect these allosteric effects. However, generating the initial high-quality lead antibodies remains a significant hurdle due to the conservation and instability of these enzymes.
Challenges in Developing Anti-Sulfotransferase Antibodies
The development of reliable antibodies against sulfotransferases presents several technical obstacles that standard protocols often fail to address:
- High Sequence Homology: Different isoforms of sulfotransferases (e.g., SULT1A1 vs. SULT1A3) share high sequence identity, making it difficult to generate antibodies that do not cross-react.
- Structural Instability: Many Golgi-resident sulfotransferases are difficult to purify in their native conformation, leading to the generation of antibodies that recognize unfolded epitopes rather than the active enzyme.
- Low Immunogenicity: As conserved intracellular or Golgi enzymes, SULTs are often tolerated by the host immune system, resulting in low-titer responses during immunization.
Streamlined Development Workflow
Inquire about Anti-ST Antibody Development
Our Service Capabilities
Creative Biolabs offers a flexible service module tailored to the needs of structural biologists and protein engineers. Whether you require a standard IgG for detection or a specific scFv sequence for mutational analysis, our platform delivers.
Custom scFv Generation for Framework Engineering
We specialize in generating anti-sulfotransferase scFv libraries. This service is specifically designed for clients investigating affinity maturation mechanisms. We can construct libraries with targeted diversity in the VH framework region 1 (FR1), allowing for the isolation of mutants with enhanced stability and binding properties without altering the CDRs. This is ideal for studying ST antibody evolution and folding kinetics.
Anti-Glycosyltransferase Specificity Screening
To ensure your antibody is specific to a sulfotransferase and not related glycosyltransferases (e.g., FUT8, ST6GAL1, or MGAT family members), we perform rigorous cross-reactivity checks. We utilize a panel of carbohydrate active enzymes to validate that the binder recognizes the unique structural motifs of the target SULT.
Recombinant Antibody Production
Once a lead anti-ST antibody is identified, we offer scalable recombinant production in HEK293 or CHO systems. We can convert scFv or Fab fragments into full-length IgG formats or fuse them with specific tags for downstream purification and crystallization studies.
Hybridoma Development
For applications requiring extremely high affinity and native epitope recognition, we employ advanced immunization strategies in mice or rabbits. This is particularly effective for generating antibodies against difficult targets like membrane-associated heparanase or polysialyltransferase.
Kinetic Analysis (SPR/BLI)
We provide detailed kinetic profiling of your anti-sulfotransferase antibodies. Using Surface Plasmon Resonance (SPR) or Bio-Layer Interferometry (BLI), we measure on-rates and off-rates, which is crucial when comparing the efficacy of different framework mutants.
Service Highlights
Defined Epitope Specificity
We generate antibodies that distinguish between closely related sulfotransferase isoforms with high precision.
Structure-Based Design
Our protocols are optimized for producing scFv candidates suitable for VH framework engineering and structural studies.
High Affinity Maturation
Access to advanced mutagenesis platforms to improve antibody KD values to the picomolar range.
Diverse Applications
Validated for Western Blot, ELISA, Flow Cytometry, and as crystallization chaperones.
How to Start
Initiating a project with Creative Biolabs is a collaborative process. We begin by defining the specific sulfotransferase target (e.g., SULT1A1, SULT1E1, or a specific heparan sulfate sulfotransferase) and the intended application. If your goal is structural engineering, we will design the scFv library generation strategy to focus on the variable heavy chain framework. For standard detection needs, we will select the most immunogenic epitopes. We provide regular updates and milestone reports, ensuring that the final product integrates seamlessly into your research workflow.
Start Your Project Now
Published Data
The expression of heparan sulfate 3-O-sulfotransferases (HS3STs) is frequently dysregulated in cancer, influencing critical cellular processes such as proliferation and invasion. A review highlights that aberrant expression of HS3STs, including HS3ST2, HS3ST3A, and HS3ST3B, has been reported in a variety of cancers such as breast, lung, and pancreatic cancer. These enzymes are key to the maturation of heparan sulfate chains, providing specific binding sites for various ligands. The schematic below illustrates the complexity of heparan sulfate modifications, emphasizing the precise role of HS3STs in the final 3-O-sulfation step (shown in red). Access to specific antibodies against these enzymes is crucial for researchers to deconvolute their pro-tumoral or anti-oncogenic roles in different tumor microenvironments.
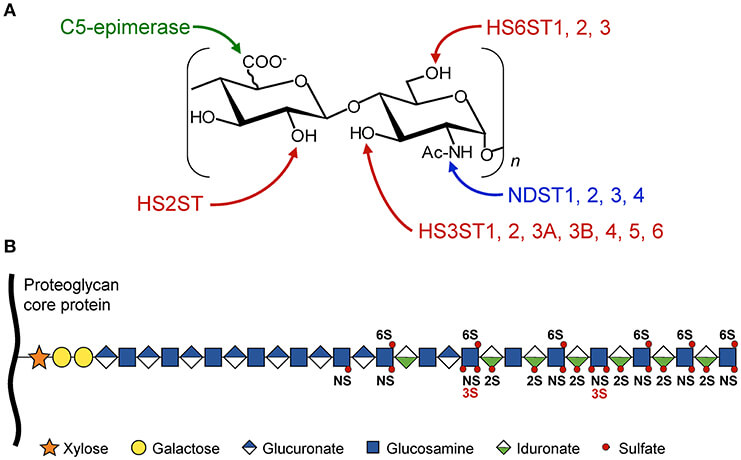 Fig.1
Schematic representation of HS modifications mediated by sulfotransferases including HS3STs.1
Fig.1
Schematic representation of HS modifications mediated by sulfotransferases including HS3STs.1
FAQs
Why is the VH framework region 1 (FR1) important for anti-sulfotransferase antibody engineering?
The VH framework region 1 plays a critical role in supporting the structural integrity of the variable domain. Research has shown that mutations in FR1 can allosterically influence the conformation of the CDR loops, thereby modulating antigen binding affinity and stability. This makes it an attractive target for affinity maturation strategies that avoid direct CDR mutagenesis.
Can you develop antibodies that distinguish between different sulfotransferase isoforms?
Yes. Sulfotransferases often share high sequence homology, but we utilize peptide-based immunization or epitope-blocking strategies during phage panning to select binders that recognize unique surface residues. We perform extensive cross-reactivity screening against related family members (e.g., distinguishing SULT1A1 from SULT1A3).
What formats of anti-ST antibodies can you provide?
We can provide antibodies in various formats including full-length IgG, Fab fragments, scFv, and VHH. For structural studies and phage display libraries, the scFv format is often preferred due to its small size and ease of genetic manipulation.
Do you offer antibodies against glycosyltransferases other than sulfotransferases?
Absolutely. Our platform covers a wide range of carbohydrate-active enzymes (CAZymes), including sialyltransferases (e.g., ST6GAL1, ST3GAL), fucosyltransferases (e.g., FUT8, FUT4), neuraminidases (NEU1, NEU3), and glycosidases.
What is the typical timeline for a custom scFv development project?
A standard phage display project for scFv generation typically takes 12-16 weeks. This includes library construction or selection, panning, screening, and preliminary validation. Projects involving complex affinity maturation or framework engineering may require additional time.
How do you validate the specificity of the anti-glycan enzyme antibodies?
Validation typically involves ELISA against the immunogen and related proteins to determine specificity. We also employ Western Blotting to check for recognition of the denatured protein and, if required, flow cytometry or immunofluorescence to verify binding to the native enzyme in a cellular context.
What Our Customers Say
"We required an anti-sulfotransferase scFv for a very specific mutational study involving the VH framework. Creative Biolabs was one of the few vendors who understood the structural requirements of our request. The library they constructed was diverse and yielded excellent candidates."
"The specificity of the antibody against our target cytosolic sulfotransferase was impressive, especially given the high homology with other isoforms. The validation data package was thorough and saved us weeks of internal testing."
Reference:
- Denys, Agnès, and Fabrice Allain. "The emerging roles of heparan sulfate 3-O-sulfotransferases in cancer." Frontiers in Oncology 9 (2019): 507. Distributed under Open Access license CC BY 4.0. https://doi.org/10.3389/fonc.2019.00507
Supports
- Glycosyltransferase & Glycosidase Substrate Microarray
- Glycosylation Analysis
- Custom Glycosylation of Biomolecules
