Sialyltransferases: Aberrant Expression in Tumorigenesis
Creative Biolabs supports glycan-focused cancer research with experience in studying sialyltransferase expression, tumor-associated hypersialylation, and glycan-related mechanisms relevant to tumor progression. For teams working on enzyme expression, target validation, reagent generation, or pathway-focused assay development, our Anti-Sialyltransferase Antibody Development service can provide research tools against selected ST family targets. We explain how aberrant sialyltransferase activity may reshape tumor-associated glycans and why these changes matter for receptor biology, cell behavior, and experimental design in oncology research.
Why Sialyltransferases Matter in Tumor Biology
Sialyltransferases catalyze the transfer of sialic acid to terminal positions on glycoproteins and glycolipids. In malignant cells, dysregulated expression of these enzymes can alter glycan density, terminal capping, receptor residence time, and ligand accessibility. The result is often a surface state described as tumor hypersialylation. In cancer biology, this change can affect cell signaling, adhesion, resistance to cell death, and immune-related interactions at the cell surface.
Among the best-studied enzymes, ST6GAL1 has received sustained attention because it installs α2,6-linked sialic acid on N-glycosylated proteins. In the literature, ST6GAL1 has been linked to receptor regulation, epithelial-to-mesenchymal transition, invasion, stem-like features, chemoresistance, resistance to cell death, and angiogenesis in selected tumor models. Other sialyltransferases, including members of the ST3GAL and ST6GALNAC families, also contribute to cancer-associated glycan remodeling by generating distinct terminal sialic acid patterns on mucins, adhesion molecules, and other glycoconjugates.
| Topic | What Changes in Cancer | Research Relevance | Typical Readout |
|---|---|---|---|
| Sialyltransferase expression | Upregulated ST genes, altered isoform balance, pathway-specific induction | Links enzyme abundance to tumor behavior and glycan phenotype | qPCR, immunoblotting, antibody staining |
| Tumor hypersialylation | Increased terminal sialic acid on glycoproteins and glycolipids | Supports studies on masking, receptor retention, and immune modulation | Lectin binding, glycomics, mass spectrometry |
| Sialic acid linkage | Shift in α2,3, α2,6, or other context-dependent terminal motifs | Helps distinguish linkage-dependent recognition and binding behavior | Linkage-sensitive probes, glycan profiling, microarray-based binding studies |
| Immune interface | Altered sialylated surface glycans that may affect immune-related interactions | Supports studies on glycan-dependent immune modulation in tumor models | Immune co-culture, binding assays, pathway analysis |
Common Biological Contexts
- Enhanced receptor stability and prolonged surface retention.
- Reduced susceptibility to death receptor-mediated apoptosis.
- Greater compatibility with invasive and stem-like tumor states.
- Potentially altered engagement of glycan-binding receptors involved in immune regulation.
- Broader remodeling of the tumor microenvironment through glycoprotein and glycolipid changes.
How Aberrant Sialyltransferase Activity Promotes Tumor Progression
The biological impact of sialyltransferase overexpression depends on which acceptor substrates are available, where the modified proteins traffic, and which ligands encounter those glycans. Even so, several recurring mechanisms appear repeatedly in the literature and are highly relevant to experimental planning.
1. Glycan Shielding of Cell Surfaces
Heavy terminal sialylation can change the accessibility of cell-surface molecules. Protein epitopes may become less exposed. Ligand presentation can shift. This matters when researchers compare tumor and normal cells, evaluate antibody binding, or interpret why strongly expressed membrane proteins do not behave as expected in immune assays.
2. Receptor Signaling Amplification
Certain glycoprotein receptors exhibit altered activation, clustering, or membrane retention after α2,6 sialylation. In this context, st6gal1 function is often discussed in connection with EGFR, integrins, TNFR1, and related pathways that support proliferation, migration, survival, or therapy resistance.
3. Apoptosis Avoidance
Sialylation can affect death receptor behavior and downstream signaling. When pro-apoptotic pathways are blunted, tumor cells gain tolerance to environmental stress and experimental perturbation. This mechanism is especially important when researchers investigate drug response, cytokine signaling, or immune effector function.
4. Immune-Related Glycan Effects
Aberrant sialylation can contribute to immune evasion in some tumor settings by changing how tumor cells interact with immune components in the microenvironment. In practice, this question usually requires direct functional validation rather than inference from glycan abundance alone.
Why This Area Is Experimentally Challenging
Work on tumor-associated sialylation often becomes difficult not because the biology is weak, but because the readouts are easy to oversimplify. Many experiments report increased sialylation without resolving which linkage increased, which substrate changed, or which immune-related effect became functionally relevant. Reliable interpretation usually requires combining enzyme-level, glycan-level, and protein-level evidence.
Linkage Complexity
Different sialic acid linkage patterns do not produce the same biological outcome. A global positive signal for sialylation is rarely enough.
Substrate Diversity
The same enzyme can modify multiple membrane proteins and secreted molecules. Functional assignment therefore needs careful validation.
Model Dependence
ST6GAL1-high findings in one tumor type may not translate directly to another because substrate pools and signaling context differ.
Immune Readout Ambiguity
Suppressed immunity can arise from multiple parallel pathways. Glycan-dependent effects need direct supporting data rather than assumption.
What Stronger Studies Usually Include
- Expression analysis of the candidate sialyltransferase at RNA and protein levels.
- Glycosylation profiling before and after perturbation.
- Linkage-sensitive characterization of terminal sialylation.
- Protein-specific evidence for modified receptors or ligands.
- Functional assays tied to migration, survival, or immune interaction.
Our Research Solutions for Sialyltransferase-Focused Projects
Creative Biolabs provides integrated research support for teams studying sialyltransferases in oncology. The most suitable combination depends on whether your main question is enzyme detection, glycan profiling, protein-level mechanism, or ligand-binding specificity.
Anti-Sialyltransferase Antibody Development
This service is the best fit for projects that require reliable reagents against ST family members, including antibody generation for expression profiling, pathway validation, cell-based assays, or custom detection workflows. It is especially useful when commercial antibodies show weak specificity, incomplete validation, or poor performance in your model system.
Glycosylation Analysis Service
Ideal for profiling broad glycan remodeling associated with malignant transformation. This service helps define whether sialyltransferase overexpression is accompanied by a measurable shift in total glycosylation patterns and provides a strong entry point for hypothesis-driven follow-up studies.
Protein Glycosylation Analysis Service
Recommended when the focus is on specific glycoproteins, such as receptors, ligands, or adhesion molecules that may gain functional changes after altered sialylation. This approach is valuable for mechanistic work on signaling persistence, membrane retention, and protein-level glycoform interpretation.
Sialoside Microarray
Useful when your study requires detailed comparison of sialylated glycan-binding preferences or interaction patterns with linkage-dependent probes, lectins, antibodies, or receptors. It is particularly valuable for distinguishing α2,3- and α2,6-dependent recognition events, although endogenous sample linkage abundance usually requires complementary analytical methods.
Suggested Service Pairings
| Research Goal | Recommended Combination | Why It Fits |
|---|---|---|
| Verify ST enzyme overexpression | Anti-Sialyltransferase Antibody Development + Glycosylation Analysis Service | Connects enzyme detection with overall glycan phenotype. |
| Study ST6GAL1 function on receptor biology | Anti-Sialyltransferase Antibody Development + Protein Glycosylation Analysis Service | Combines target validation with protein-specific glycoform evidence. |
| Resolve sialic acid linkage effects | Sialoside Microarray + Protein Glycosylation Analysis Service | Connects binding preference data with protein-level glycosylation context. |
| Build a broader tumor hypersialylation workflow | Anti-Sialyltransferase Antibody Development + Glycosylation Analysis Service + Protein Glycosylation Analysis Service + Sialoside Microarray | Supports enzyme, glycan, protein, and binding analyses in one pipeline. |
Talk with Our Team About Your Project
If your group is investigating sialyltransferases in tumorigenesis, the most useful project briefing usually includes the cancer model, the target enzyme or enzyme family, the preferred sample type, the main biological hypothesis, and the downstream assay format. That information helps define whether your study needs a custom antibody, whole-glycome profiling, protein-centered glycosylation analysis, or linkage-focused microarray work.
Our team can help you design a research workflow around expression confirmation, linkage analysis, protein-level glycan characterization, or glycan interaction studies. Share your target, sample type, and analytical priority through the online inquiry form to receive a tailored recommendation.
Request a Project Discussion
Published Data
A 2022 open-access review in Frontiers in Molecular Biosciences summarized how ST6Gal1-mediated α2,6 sialylation relates to multiple oncogenic pathways and cancer hallmarks. The authors describe ST6Gal1 as a Golgi sialyltransferase that adds α2,6-linked sialic acid to terminal galactose on N-glycoproteins and note that elevated ST6Gal1 expression has been reported in multiple malignancies. The review discusses links between ST6Gal1-associated signaling and sustained proliferation, enhanced self-renewal, epithelial-to-mesenchymal transition, invasion, chemoresistance, resistance to cell death, angiogenesis, and immune evasion, while also emphasizing that many mechanisms still need further delineation.
The figure integrates receptor-level examples and phenotype-level consequences in one scheme. It shows membrane-bound N-glycoproteins discussed in the review, including Fas, TNFR1, β1-integrin, PECAM, VEGFR2, and EGFR, together with pathways linked to TGF-β, PI3K/AKT, GSK3β, β-catenin, and Wnt. In the schematic, ST6Gal1-associated α2,6 sialylation is presented as influencing surface retention, stability, clustering, activation, and downstream phenotypes such as survival, invasion/metastasis, stemness, and drug resistance.
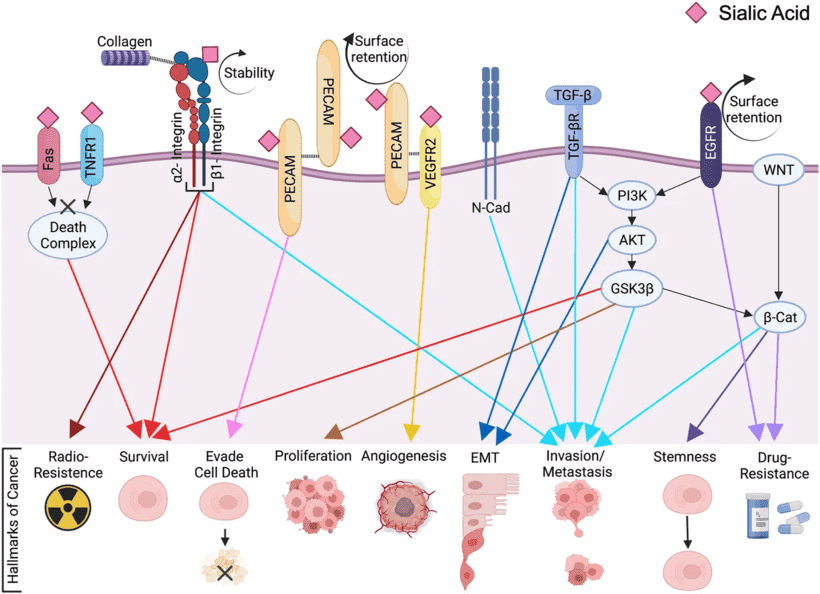 Fig.1 ST6Gal1-mediated signaling pathways in cancer progression.1
Fig.1 ST6Gal1-mediated signaling pathways in cancer progression.1
This literature is particularly helpful for framing hypothesis generation. It does not claim that every pathway is universally dominant in every tumor model. Instead, it offers a structured summary of reported ST6Gal1-related pathways and targets that can inform glycosylation analysis, antibody validation, and binding studies in cancer research.
FAQs
What is the difference between high sialyltransferase expression and tumor hypersialylation?
High sialyltransferase expression refers to increased abundance of one or more enzymes. Tumor hypersialylation refers to the resulting glycan phenotype on tumor-associated proteins or lipids. The two are related, but they are not interchangeable because substrate availability and pathway context also shape the final glycan output.
Why is ST6GAL1 discussed so often in cancer glycosylation studies?
ST6GAL1 is one of the most studied sialyltransferases because it installs α2,6-linked sialic acid on N-glycoproteins and has been associated with receptor regulation, invasion, survival, stem-like traits, and therapy resistance across multiple tumor types.
How can I determine whether α2,3 or α2,6 sialic acid linkage is more relevant in my model?
A linkage-focused strategy is usually required. Linkage-sensitive probes, glycan profiling, and targeted glycosylation studies are usually more informative than a single total sialylation readout. Sialoside microarray can further help compare binding preferences toward different terminal motifs.
Can glycosylation analysis alone explain cancer immune evasion?
Not by itself. Glycosylation analysis can show whether glycan remodeling is present, but functional immune interpretation usually also requires receptor-level evidence, binding assays, or immune co-culture data. Strong conclusions come from combining glycan profiling with biological validation.
When should I choose custom anti-sialyltransferase antibodies instead of standard catalog reagents?
Custom development is especially useful when catalog antibodies show weak specificity, limited application validation, poor cross-reactivity profiles, or inadequate performance in your tumor model, sample preparation method, or assay platform.
Is protein glycosylation analysis necessary if I already know the enzyme is overexpressed?
It is often necessary when the biological question involves mechanism. Enzyme overexpression suggests potential activity, but protein-level glycosylation analysis helps identify which receptors or ligands are actually remodeled and therefore more likely to drive the observed phenotype.
Are these services suitable for clinical diagnosis or treatment decisions?
No. The services described on this page are provided for research use only. They support basic, translational, and preclinical investigation of glycan biology but are not intended for clinical diagnosis or therapeutic use.
Reference:
- Sajina, S. G. C., Susan L. Bellis, and Anita B. Hjelmeland. ST6Gal1: Oncogenic signaling pathways and targets. Frontiers in Molecular Biosciences 9 (2022): 962908. Distributed under Open Access license CC BY 4.0, without modification. https://doi.org/10.3389/fmolb.2022.962908
