Glyco-Enzymes: The Builders and Trimmers of the Glycome
Glyco-enzymes determine how glycans are built, edited, and recycled across cells, tissues, and experimental systems. For research teams studying glycan architecture, enzyme inhibition, or pathway-specific remodeling, Creative Biolabs offers dedicated anti-glycan related enzyme antibody development services to support target validation, detection workflows, and glycobiology tool development. In practice, understanding glyco-enzyme function requires looking at both sides of the pathway: glycosyltransferases that install sugars and glycosidases that trim, remodel, or degrade them. Their coordinated action underlies glycome biosynthesis, glycoprotein maturation, glycolipid processing, and glycan turnover in research models.
Why Glyco-Enzymes Matter in Glycobiology
Glycans are not assembled from a direct template in the way nucleic acids or proteins are. Instead, they arise from enzyme-regulated pathways distributed across the endoplasmic reticulum, Golgi apparatus, cell surface, extracellular space, and lysosome. That is why glyco-enzymes sit at the center of modern glycobiology.
Spatial Control
Distinct enzyme sets occupy defined subcellular compartments, so glycan composition depends on where a substrate travels and how long it remains accessible.
Substrate Competition
Multiple enzymes may compete for the same acceptor or donor pool, creating branch points that reshape glycan complexity and distribution.
Dynamic Remodeling
Trimming and reprocessing steps alter immature glycans into mature structures, and post-secretory glycosidases continue remodeling after export.
Functional Consequences
Enzyme activity influences folding, trafficking, receptor binding, matrix interactions, and the analytical readout of many glycobiology experiments.
Because glycans are generated by pathway logic rather than a one-gene-one-product rule, enzyme-level analysis often provides the clearest route to mechanistic insight. Researchers frequently begin with a glycan phenotype, but the decisive questions are usually enzymatic: which transferase added the residue, which glycosidase removed it, which donor pool was limiting, and which perturbation is most informative.
Glycosyltransferase vs Glycosidase: A Practical Comparison
The simplest way to frame glycosyltransferase vs glycosidase biology is to view one group as builders and the other as trimmers. That distinction is useful, but real systems are more nuanced because both enzyme classes can promote maturation, quality control, or turnover depending on context.
| Feature | Glycosyltransferases | Glycosidases |
|---|---|---|
| Primary reaction | Transfer activated monosaccharides from donor molecules to proteins, lipids, or growing glycans | Hydrolyze glycosidic bonds to trim, remodel, or degrade glycan structures |
| Main role in pathways | Chain initiation, elongation, branching, capping, and linkage-specific diversification | Processing of immature intermediates, quality control, terminal editing, and lysosomal turnover |
| Common locations | ER and Golgi, with additional roles in cytosol and specific organelles | ER, Golgi, lysosome, cell surface, and extracellular matrix |
| Representative examples | ALG enzymes, OST complex, MGAT family, FUTs, STs, B4GALTs, GALNTs, OGT | Alpha-mannosidases, glucosidases, neuraminidases, lysosomal exoglycosidases, OGA |
| Typical experimental questions | Which enzyme installs a residue, creates branching, or generates a target epitope? | Which enzyme removes a residue, edits maturation, or controls glycan turnover? |
How the two enzyme classes cooperate
- In N-glycosylation, early transferases assemble precursor glycans, while ER and Golgi glucosidases and mannosidases trim intermediates before later transferases extend them.
- In mucin-type O-glycosylation, initiation by GalNAc-transferases is followed by extension and capping through additional transferases, while selective glycosidase activity can reshape terminal motifs.
- In O-GlcNAc cycling, OGT adds GlcNAc to nuclear and cytoplasmic proteins, whereas OGA removes it, creating a reversible regulatory system rather than one-way biosynthesis.
- In lysosomal pathways, glycosidases release monosaccharides from glycoconjugates, enabling recycling that indirectly feeds future glycome biosynthesis.
Core Steps That Shape Glycome Biosynthesis
Glycome biosynthesis depends on enzyme sequence, compartmentalization, donor supply, and substrate accessibility. For most research programs, the most informative framework is to divide the process into four operational layers.
1. Donor Sugar Generation
Activated sugar donors such as UDP-GlcNAc, GDP-Fuc, CMP-Sia, and UDP-Gal are generated from central metabolic pathways. Their availability influences branching, terminal decoration, and pathway throughput. This is why metabolism and glycosylation are tightly linked.
2. Compartment-Specific Processing
Substrates move through the ER and Golgi where distinct enzyme sets act in sequence. Early transferases install core structures. Glycosidases remove selected residues. Later transferases generate branching, elongation, fucosylation, galactosylation, sialylation, or specialized caps.
3. Surface and Matrix Remodeling
After secretion or membrane delivery, glycans may still be edited by extracellular or membrane-associated glycosidases. These events can change receptor availability, ligand recognition, and local glycan exposure in a way that matters for cell-based assays.
4. Turnover and Recycling
Lysosomal glycosidases break down glycoconjugates into reusable monosaccharides. This is not only a degradative endpoint. It also supports cellular economy and helps maintain the long-term balance of the glycan system.
Major pathway contexts researchers usually evaluate
| Pathway Context | Key Enzyme Logic | Common Research Readouts |
|---|---|---|
| N-glycan maturation | Core assembly, glucose and mannose trimming, branching, fucosylation, galactosylation, sialylation | Site occupancy, glycoform distribution, trafficking effects, glycoprotein quality |
| Mucin-type O-glycans | Initiation by GalNAc transferases followed by core extension and terminal capping | Epitope exposure, mucin profiling, cell-surface glycan characterization |
| Glycolipid biosynthesis | Sequential transferase steps generate glycosphingolipid series and terminal motifs | Ganglioside mapping, membrane phenotype studies, antibody target discovery |
| O-GlcNAc cycling | Reversible addition by OGT and removal by OGA | Signal transduction, stress response, nutrient sensing, proteomic profiling |
| Lysosomal glycan turnover | Stepwise cleavage by exoglycosidases and related hydrolases | Turnover analysis, catabolic bottlenecks, substrate accumulation studies |
What Controls Glyco-Enzymes Function
Even when gene annotations are known, enzyme output can vary widely across systems. That variation is one reason glycobiology tools must be selected carefully and validated with pathway context in mind.
- Subcellular localization and Golgi positioning
- Expression level and isoenzyme redundancy
- Donor substrate availability and metabolic flux
- Acceptor accessibility on proteins, lipids, or nascent glycans
- Competition among enzymes for shared substrates
- Complex formation, trafficking, and retention mechanisms
- pH, cofactors, and assay-specific reaction conditions
These factors explain why an enzyme knockout, overexpression model, inhibitor study, antibody readout, or glycan array result cannot be interpreted in isolation. In many projects, the most rigorous approach combines enzyme-specific detection with glycan-level phenotyping.
Research Uses: From Enzyme Inhibition to Glycobiology Tools
For discovery teams, glyco-enzymes are not only pathway components. They are also practical intervention points and analytical anchors. A focused study design may start from a single enzyme, a glycan motif, or a phenotype that changes after perturbation.
Target Validation
Enzyme-specific antibodies, expression profiling, and orthogonal glycan measurements help confirm whether a candidate transferase or glycosidase truly drives the observed glycan phenotype.
Pathway Mapping
Sequential perturbation strategies can separate early processing defects from later branching or terminal capping events and clarify the order of pathway dependencies.
Enzyme Inhibition Studies
Small molecules, genetic perturbations, and substrate competition approaches are often used to test how inhibition reshapes glycan output rather than simply whether an enzyme is present.
Antibody and Binder Discovery
Antibodies against glycan-related enzymes support western blotting, IHC, IF, flow cytometry, and mechanistic studies where enzyme localization is as informative as abundance.
Cell Model Engineering
Engineered lines with altered transferase or glycosidase activities provide defined glycan backgrounds for ligand binding, antibody validation, and comparative glycomics.
Analytical Workflow Support
Microarrays, glycosylation analysis, and binding assays help convert enzyme hypotheses into experimentally testable pathway models.
Our Solutions for Glycan-Related Enzyme Research
Creative Biolabs supports enzyme-focused glycoscience projects with flexible service combinations. Whether your starting point is a glycosyltransferase, a glycosidase, or a downstream glycan phenotype, we can align detection reagents and analytical tools to the biological question.
- Anti-Glycan Related Enzyme Antibody Development for customized binders against glycan-processing targets
- Glycoarray Platforms for interaction profiling and glycan-focused validation
- Glycosylation Analysis for glycoproteins, antibodies, glycolipids, and related biomolecules
- Surface Plasmon Resonance for affinity and kinetics characterization
What we can help you evaluate
Our project teams routinely support questions such as enzyme expression pattern, reagent specificity, pathway-dependent glycan changes, target suitability for enzyme inhibition studies, and assay compatibility across different sample types. We can also help design validation workflows that connect enzyme-level readouts with glycan structural evidence instead of treating them as separate data streams.
Custom Antibodies for Glycan-Related Enzymes
We develop research antibodies against glycosyltransferases, glycosidases, and associated pathway proteins with attention to target class, localization, sequence conservation, and downstream assay needs.
Substrate and Specificity Profiling
For teams studying glyco-enzymes function, substrate scope, or selectivity, our platform options support higher-confidence interpretation of activity and binding behavior.
Start with a pathway question, not only a target list
When you contact us, the most useful information includes the enzyme class, target species, sample type, intended applications, expected glycan context, and whether you are prioritizing localization, pathway perturbation, or direct assay development. That makes it easier to recommend a workflow that fits your actual research endpoint.
Discuss Your Glyco-Enzyme Research Project
Published Data
A 2024 open access mini-review on metabolism-driven glycosylation in interstitial lung diseases includes a pathway schematic that is useful for enzyme-focused glycobiology research. In that figure, the authors connect the hexosamine biosynthetic pathway with ER-Golgi protein glycosylation, O-GlcNAc cycling, extracellular glycosidase activity, and lysosomal glycan turnover. The figure also shows activated sugar donors such as UDP-GlcNAc, GDP-Fuc, CMP-Sia, UDP-GalNAc, and UDP-Glc as inputs for glycosylation pathways. Used in this limited sense, the paper provides a helpful visual summary of how donor generation, biosynthetic transfer, trimming, and recycling are linked at the pathway level.
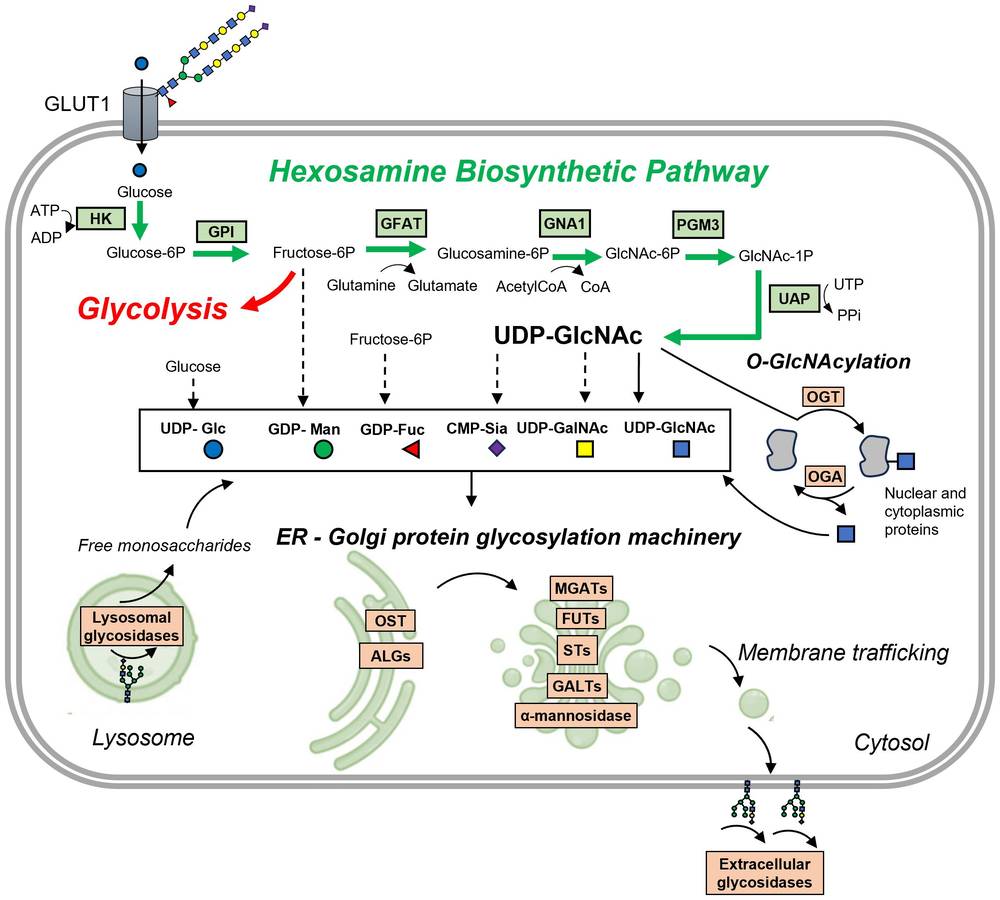 Fig. 1 Pathway view linking the hexosamine biosynthetic pathway, ER-Golgi glycosylation, O-GlcNAc cycling, and glycan turnover.1
Fig. 1 Pathway view linking the hexosamine biosynthetic pathway, ER-Golgi glycosylation, O-GlcNAc cycling, and glycan turnover.1
The same review also notes that glycosylation mainly proceeds through ER and Golgi machinery, while additional remodeling can occur through extracellular glycosidases and lysosomal glycosidases. It further explains that UDP-GlcNAc, produced through the hexosamine biosynthetic pathway, is a key metabolic input for glycan biosynthesis and can influence branching through enzymes such as the MGAT family. Because the article is centered on macrophages and interstitial lung diseases rather than broad method development, the value of this citation here is primarily its pathway summary and the mechanistic link it draws between metabolism and glycosylation.
FAQs
What is the main difference between glycosyltransferase and glycosidase activity?
Glycosyltransferases add sugars from activated donor molecules to acceptors such as proteins, lipids, or growing glycans, whereas glycosidases cleave glycosidic bonds to trim, remodel, or degrade glycans.
Why are both enzyme classes needed for glycome biosynthesis?
The glycome is produced through sequential assembly and controlled editing. Transferases build core and terminal structures, while glycosidases remove selected residues to enable maturation, quality control, remodeling, and turnover.
Does enzyme expression alone predict the final glycan structure?
Not reliably. Final glycan output also depends on donor availability, enzyme localization, substrate accessibility, competing enzymes, trafficking, and the time a substrate spends in each compartment.
How is enzyme inhibition usually interpreted in glycobiology studies?
The most informative readout is usually the downstream change in glycan structure or abundance, not only reduced target signal. Enzyme inhibition studies are strongest when paired with glycan-level validation.
Can glycosidases also regulate mature glycans outside the Golgi?
Yes. Certain glycosidases act at the cell surface, in the extracellular matrix, or in lysosomes, where they contribute to remodeling, turnover, and monosaccharide recycling.
What kinds of reagents are useful for studying glycan-related enzymes?
Common glycobiology tools include enzyme-specific antibodies, glycan arrays, glycosylation analysis platforms, engineered cell models, and binding assays that connect pathway changes to structure-level evidence.
What information should I provide when requesting a glyco-enzyme research service?
Useful starting details include the target enzyme, species, sample type, preferred application, expected glycan context, and whether your main goal is detection, pathway mapping, or assay development.
Reference:
- Drzewicka, Katarzyna, and Zbigniew Zasłona. "Metabolism-driven glycosylation represents therapeutic opportunities in interstitial lung diseases." Frontiers in Immunology 15 (2024): 1328781. Distributed under Open Access license CC BY 4.0, without modification. https://doi.org/10.3389/fimmu.2024.1328781
